Abstract
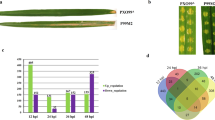
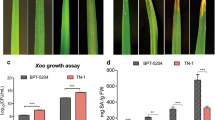
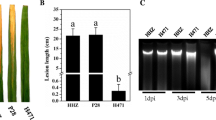
Xanthomonas oryzae pv. oryzae (Xoo) is a destructive pathogen that causes bacterial blight disease of rice worldwide. Xoo uses T3SS (type III secretion system) effectors to subvert rice innate immunity. However, the comprehensive knowledge of rice genes involved in T3SS effectors-mediated interaction remains unclear. In this study, the transcriptome profiles of rice infected with a virulent Xoo strain from North-eastern region of India relatives to its avirulent strain (that lacks functional T3SS) were analyzed at early (2–6 hpi) and late (16–24 hpi) hours of infection. Out of total 255 differentially expressed genes (DEGs), during early infection, 62 and 70 genes were upregulated and downregulated, respectively. At late infection, 70 and 53 genes were upregulated and downregulated, respectively. The transcriptomic data identified many differentially expressed resistant genes, transposons, transcription factors, serine/threonine protein kinase, cytochrome P450 and peroxidase genes that are involved in plant defense. Pathway analysis revealed that these DEGs are involved in hormone signaling, plant defense, cellular metabolism, growth and development processes. DEGs associated with plant defense were also validated through quantitative real-time PCR. Our study brings a comprehensive picture of the rice genes that are being differentially expressed during bacterial blight infection. Nevertheless, the DEG-associated pathways would provide sensible targets for developing resistance to bacterial blight.






Similar content being viewed by others
References
Almagro L, Gómez Ros LV, Belchi-Navarro S, Bru R, Ros Barceló A, Pedreño MA (2009) Class III peroxidases in plant defence reactions. J Exp Bot 60:377–390. https://doi.org/10.1093/jxb/ern277
Bittner-Eddy PD, Crute IR, Holub EB, Betnon JL (2000) RPP13 is a simple locus in Arabidopsis thaliana for alleles that specify downy mildew resistance to different avirulence determinants in Peronospora parasitica. Plant J 21:177–188. https://doi.org/10.1046/j.1365-313x.2000.00664.x
Cesari S, Thilliez G, Ribot C, Chalvon V, Michel C, Jauneau A, Rivas S, Alaux L, Kanzaki H, Okuyama Y, Morel JB (2013) The rice resistance protein pair RGA4/RGA5 recognizes the Magnaporthe oryzae effectors AVR-Pia and AVR1-CO39 by direct binding. Plant Cell 25:1463–1481. https://doi.org/10.1105/tpc.112.107201
Cheema KK, Grewal NK, Vikal Y, Sharma R, Lore JS, Das A, Bhatia D, Mahajan R, Gupta V, Bharaj TS, Singh K (2008) A novel bacterial blight resistance gene from Oryza nivara mapped to 38 kb region on chromosome 4L and transferred to Oryza sativa L. Genet Res (camb) 90:397–407. https://doi.org/10.1017/S0016672308009786
Christie N, Tobias PA, Naidoo S, Külheim C (2016) The Eucalyptus grandis NBS-LRR gene family: physical clustering and expression hotspots. Front Plant Sci 6:1238. https://doi.org/10.3389/fpls.2015.01238
Cox KL, Meng F, Wilkins KE, Li F, Wang P, Booher NJ, Carpenter SC, Chen LQ, Zheng H, Gao X, Zheng Y (2017) TAL effector driven induction of a SWEET gene confers susceptibility to bacterial blight of cotton. Nat Commu 8:15588. https://doi.org/10.1038/ncomms15588
DeYoung BJ, Innes RW (2006) Plant NBS-LRR proteins in pathogen sensing and host defense. Nat Immunol 7:1243–1249. https://doi.org/10.1038/ni1410
Ellur RK, Khanna A, Gopala Krishnan S, Bhowmick PK, Vinod KK, Nagarajan M, Mondal KK, Singh NK, Prabhu KV, Singh AK (2016a) Marker-aided incorporation of Xa38, a novel bacterial blight resistance gene, in PB1121 and comparison of its resistance spectrum with xa13 + Xa21. Sci Rep 6:29188. https://doi.org/10.1038/srep29188
Ellur RK, Khanna A, Yadav A, Pathania S, Rajashekara H, Singh VK, Krishnan SG, Bhowmick PK, Nagarajan M, Vinod KK, Prakash G (2016b) Improvement of Basmati rice varieties for resistance to blast and bacterial blight diseases using marker assisted backcross breeding. Plant Sci 242:330–341. https://doi.org/10.1016/j.plantsci.2015.08.020
Harkenrider M, Sharma R, De Vleesschauwer D, Tsao L, Zhang X, Chern M, Canlas P, Zuo S, Ronald PC (2016) Overexpression of rice wall-associated kinase 25 (OsWAK25) alters resistance to bacterial and fungal pathogens. PLoS ONE 11:e0147310. https://doi.org/10.1371/journal.pone.0147310
Huot B, Yao J, Montgomery BL, He SY (2014) Growth–defense tradeoffs in plants: a balancing act to optimize fitness. Mol Plant 7:1267–1287. https://doi.org/10.1093/mp/ssu049
Hutin M, Pérez-Quintero AL, Lopez C, Szurek B (2015) MorTAL Kombat: the story of defense against TAL effectors through loss-of-susceptibility. Front Plant Sci 6:535. https://doi.org/10.3389/fpls.2015.00535
Kim SM, Suh JP, Qin Y, Noh TH, Reinke RF, Jena KK (2015) Identification and fine-mapping of a new resistance gene, Xa40, conferring resistance to bacterial blight races in rice (Oryza sativa L.). Theor Appl Genet 128:1933–1943. https://doi.org/10.1007/s00122-015-2557-2
Lee H, Cha J, Choi C, Choi N, Ji HS, Park SR, Lee S, Hwang DJ (2018) Rice WRKY11 plays a role in pathogen defense and drought tolerance. Rice 11:5. https://doi.org/10.1186/s12284-018-0199-0
Li Z, Huang J, Wang Z, Meng F, Zhang S, Wu X, Zhang Z, Gao Z (2019) Overexpression of Arabidopsis nucleotide-binding and leucine-rich repeat genes RPS2 and RPM1 (D505V) confers broad-spectrum disease resistance in rice. Front Plant Sci 10:417. https://doi.org/10.3389/fpls.2019.00417
Li Q, Hu A, Qi J, Dou W, Qin X, Zou X, Xu L, Chen S, He Y (2020) CsWAKL08, a pathogen-induced wall-associated receptor-like kinase in sweet orange, confers resistance to citrus bacterial canker via ROS control and JA signaling. Hortic Res 7:42. https://doi.org/10.1038/s41438-020-0263-y
Liu L, Mei Q, Yu Z, Sun T, Zhang Z, Chen M (2013) An integrative bioinformatics framework for genome-scale multiple level network reconstruction of rice. J Integr Bioinform 10:94–102. https://doi.org/10.2390/biecoll-jib-2013-223
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Maeda K, Houjyou Y, Komatsu T, Hori H, Kodaira T, Ishikawa A (2009) AGB1 and PMR5 contribute to PEN2-mediated preinvasion resistance to Magnaporthe oryzae in Arabidopsis thaliana. Mol Plant Microbe Interact 22:1331–1340. https://doi.org/10.1094/MPMI-22-11-1331
Mondal KK (2016) Emerging phytobacterial diseases in india: research status and challenges. In: Perspectives of plant pathology in genomic era. India Phytopathological Society, New Delhi, pp 167–178
Mondal KK, Meena BR, Junaid A, Verma G, Mani C, Majumder D, Khicher M, Kumar S, Banik S (2014) Pathotyping and genetic screening of type III effectors in Indian strains of Xanthomonas oryzae pv. oryzae causing bacterial leaf blight of rice. Physiol Mol Plant Pathol 86:98–106. https://doi.org/10.1016/j.pmpp.2014.03.005
Mondal KK, Verma G, Manju JA, Mani C (2016) Rice pathogen Xanthomonas oryzae pv. oryzae employs inducible hrp-dependent XopF type III effector protein for its growth, pathogenicity and for suppression of PTI response to induce blight disease. Eur J Plant Pathol 14:311–323. https://doi.org/10.1007/s10658-015-0768-7
Naithani S, Gupta P, Preece J, D’Eustachio P, Elser JL, Garg P, Dikeman DA, Kiff J, Cook J, Olson A, Wei S (2020) Plant Reactome: a knowledgebase and resource for comparative pathway analysis. Nucleic Acids Res 48:D1093–D1103. https://doi.org/10.1093/nar/gkz996
Okada A, Shimizu T, Okada K, Kuzuyama T, Koga J, Shibuya N, Nojiri H, Yamane H (2007) Elicitor induced activation of the methylerythritol phosphate pathway toward phytoalexins biosynthesis in rice. Plant Mol Biol 65:177–187. https://doi.org/10.1007/s11103-Mol.Plant-MicrobeInteract.007-9207-2
Park HL, Bhoo SH, Kwon M, Lee SW, Cho MH (2017) Biochemical and expression analyses of the rice cinnamoyl-CoA reductase gene family. Front Plant Sci 8:2099. https://doi.org/10.3389/fpls.2017.02099
Sasaki K, Iwai T, Hiraga S, Kuroda K, Seo S, Mitsuhara I, Miyasaka A, Iwano M, Ito H, Matsui H, Ohashi Y (2004) Ten rice peroxidases redundantly respond to multiple stresses including infection with rice blast fungus. Plant Cell Physiol 45:1442–1452. https://doi.org/10.1093/pcp/pch165
Sato Y, Namiki N, Takehisa H, Kamatsuki K, Minami H, Ikawa H, Ohyanagi H, Sugimoto K, Itoh JI, Antonio BA, Nagamura Y (2013) RiceFREND: a platform for retrieving coexpressed gene networks in rice. Nucleic Acids Res 41:D1214–D1221. https://doi.org/10.1093/nar/gks1122
Sekhwal MK, Li P, Lam I, Wang X, Cloutier S, You FM (2015) Disease resistance gene analogs (RGAs) in plants. Int J Mol Sci 16:19248–19290. https://doi.org/10.3390/ijms160819248
Tian D, Wang J, Zeng X, Gu K, Qiu C, Yang X, Zhou Z, Goh M, Luo Y, Murata-Hori M, White FF (2014) The rice TAL effector-dependent resistance protein XA10 triggers cell death and calcium depletion in the endoplasmic reticulum. Plant Cell 26:497–515. https://doi.org/10.1105/tpc.113.119255
Van Ooijen G, Mayr G, Kasiem MM, Albrecht M, Cornelissen BJ, Takken FL (2008) Structure-function analysis of the NB-ARC domain of plant disease resistance proteins. J Exp Bot 59:1383–1397. https://doi.org/10.1093/jxb/ern045
Velazhahan R, Jayaraj J, Khabbaz S, Kumar RS, Muthukrishnan S (2006) Xanthomonas oryzae pv. oryzae infection triggers accumulation of phenolics, defense-related enzymes and thaumatin-like proteins in rice leaves. Arch Phytopathol Plant Prot 39:329–339. https://doi.org/10.1080/03235400500222172
Verma G, Sharma M, Mondal KK (2018) XopR TTSS-effector regulates in planta growth, virulence of Indian strain of Xanthomonas oryzae pv. oryzae via suppressing reactive oxygen species production and cell wall-associated rice immune responses during blight induction. Funct Plant Biol 45:561–574. https://doi.org/10.1071/FP17147
Verma G, Mondal KK, Kulshreshtha A, Sharma M (2019) XopR T3SS-effector of Xanthomonas oryzae pv. oryzae suppresses cell death-mediated plant defense response during bacterial blight development in rice. 3 Biotech 9:272. https://doi.org/10.1007/s13205-019-1802-9
White FF, Yang B (2009) Host and pathogen factors control-ling the rice-Xanthomonas oryzae interaction. Plant Physiol 150:1677–1686. https://doi.org/10.1104/pp.109.139360
White FF, Potnis N, Jones JB, Koebnik R (2009) The type III effectors of Xanthomonas. Mol Plant Pathol 10:749–766. https://doi.org/10.1111/j.1364-3703.2009.00590.x
Yang B, Sugio A, White FF (2006) Os8N3 is a host disease-susceptibility gene for bacterial blight of rice. Proc Natl Acad Sci USA 103:10503–10508. https://doi.org/10.1073/pnas.0604088103
Acknowledgements
The authors acknowledge the Director, Joint Director (Research), Head (Division of Plant Pathology), ICAR-Indian Agricultural Research Institute, New Delhi, India for their support whenever required during the execution of the project. The authors are also grateful to the scientific team of AgriGenome Labs Pvt Ltd, Smart City Kochi, Kerala, India for their support in generating the transcriptome data.
Funding
This research was funded by Department of Biotechnology, Government of India, grant number BT/PR16238/NER/95/103/2015.
Author information
Authors and Affiliations
Contributions
Conceptualization: KKM, PJH; methodology: AK, YR, GV, AB, AQ, AL, KR, MS, TG, R, ER, SM, K, NS, CM; Formal analysis and investigation: AK, KKM, DB; writing—original draft preparation: KKM, AK; writing—review and editing: KKM, AK; Funding acquisition: KKM, PJH; resources: KKM, PJH; supervision: KKM, PJH.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Research involving human participants and/or animals
This research does not include any human or animal participants.
Informed consent
This research does not include any human or animal participants.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Mondal, K.K., Kulshreshtha, A., Handique, P.J. et al. Rice transcriptome upon infection with Xanthomonas oryzae pv. oryzae relative to its avirulent T3SS-defective strain exposed modulation of many stress responsive genes. 3 Biotech 12, 130 (2022). https://doi.org/10.1007/s13205-022-03193-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03193-4