Summary
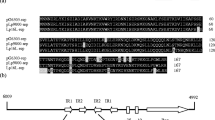
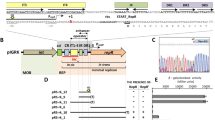
The structure of a 1.5-kb DNA sequence that is necessary and sufficient for the replication of an 8.2-kb cryptic plasmid, pFTB14, isolated from a strain of Bacillus amyloliquefaciens has been characterized. The 1.5-kb DNA sequence contains an open reading frame, rep, stretching for 1017 bp, a promoter region for rep expression, and a possible replication origin for the plasmid upstream of the promoter. The rep product is trans-active and essential for plasmid replication. The predicted rep protein is a basic protein, as are the RepC protein of pT181, RepB of pUB110 and protein A of pC194 (all these found in staphylococci) and the π protein of the R6K plasmid of Escherichia coli. The predicted rep protein has highly homologous amino acid sequences with protein A of pC194 and RepC of pUB110 throughout the protein molecule, but not with RepC of pT181, π of R6K or protein RepH encoded by and iniating the replication of pC194.
Similar content being viewed by others
References
Anagnostopoulos C, Spizizen J (1961) Requirements for transformation in Bacillus subtilis. J Bacteriol 81:741–746
Ano T, Imanaka T, Aiba S (1986) The copy number of Bacillus plasmids pRBH1 is negatively controlled by RepB protein. Mol Gen Genet 202:416–420
Bachmann BJ (1983) Linkage map of Escherichia coli K-12, Edition 7. Microbiol Rev 47:180–230
Bolivar F, Rodriguez RL, Betlach MC, Boyer HW (1977a) Construction and characterization of new cloning vehicles: I Ampicillin-resistant derivatives of the plasmid pMB9. Gene 2:75–93
Bolivar F, Rodriguez RL, Greene PJ, Betlach MC, Heyneker HL, Boyer HW, Crosa JH, Falkow S (1977b) Construction and characterization of new cloning vehicles: II A multipurpose cloning system. Gene 2:95–113
Casadaban MJ, Marinez-Arias A, Shapira SK, Chou J (1983) β-Galactosidase gene fusions for analyzing gene expression in Escherichia coli and yeast. Methods Enzymol 100:293–308
Chang S, Cohen SN (1979) High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet 168:111–115
Clarke L, Carbon J (1978) Functional expression of cloned yeast DNA in Escherichia coli: specific complementation of argininosuccinate lyase (argH) mutations. J Mol Biol 120:517–532
Donnelly CE, Sonenshein AL (1982) Genetic fusion of E. coli lac genes to a B. subtilis promoter. In: Ganesan AT, Chang S, Hoch JA (eds) Molecular cloning and gene regulation in bacilli. Academic Press, New York, pp 63–72
Germino J, Bastia D (1982) Primary structure of the replication initiation protein of plasmid R6K. Proc Natl Acad Sci USA 79:5475–5479
Horinouchi S, Weisblum B (1982a) Nucleotide sequence and functional map of pE194, a plasmid that specifies inducible resistance to macrolide, lincosamide, and streptogramin type B antibiotics. J Bacteriol 150:804–814
Horinouchi S, Weisblum B (1982b) Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J Bacteriol 150:815–825
Iordanescu S, Surdeanu M (1983) Isolation and complementation of temperature-sensitive replication mutants of Staphylococcus aureus plasmid pC194. Mol Gen Genet 191:201–206
Khan SA, Adler GK, Novick RP (1982) Functional origin of replication of pT181 plasmid DNA is contained within a 168-basepair segment. Proc Natl Acad Sci USA 79:4580–4584
Khan SA, Novick RP (1983) Complete nucleotide sequence of pT181, a tetracycline-resistance plasmid from Staphylococcus aureus. Plasmid 10:251–259
Koepsel RR, Murray RW, Rosenblum WD, Khan SA (1985) Purification of pT181-encoded repC protein required for the initiation of plasmid replication. J Biol Chem 260:8571–8577
Kumar CC, Novick RP (1985) Plasmid pT181 replication is regulated by two countertranscripts. Proc Natl Acad Sci USA 82:638–642
Legerski RJ, Hodnett JL, Gray HB, Jr (1978) Extracellular nucleases of Pseudomonas BAL31. III Use of the double-strand deoxyriboexonuclease activity as the basis of a convenient method for the mapping of fragments of DNA produced by cleavage with restriction enzymes. Nucleic Acids Res 5:1445–1464
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Maxam AM, Gilbert W (1980) Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol 65:499–560
McLaughlin JR, Murray CL, Rabinowitz JC (1981) Unique features in the ribosome binding site sequence of the gram-positive Staphylococcus aureus β-lactamase gene. J Biol Chem 256:11283–11291
Miller JH (1972) Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York, pp 352–359
Moran CP, Jr, Lang N, LeGrice SFJ, Lee G, Stephans M, Sonenshein AL, Pero J, Losick R (1982) Nucleotide sequence that signal the initiation of transcription and translation in Bacillus subtilis. Mol Gen Genet 186:339–346
Morrison DA (1977) Transformation in Escherichia coli: cryogenic preservation of competent cells. J Bacteriol 132:349–351
Muller RE, Ano T, Imanaka T, Aiba S (1986) Complete nucleotide sequences of Bacillus plasmids pUB110dB, pRBH1 and its copy mutants. Mol Gen Genet 202:169–171
Novick RP, Adler GK, Majumder S, Khan SA, Carleton S, Rosenblum WD, Iordanescu S (1982) Coding sequence for the pT181 repC product: a plasmid-coded protein uniquely required for replication. Proc Natl Acad Sci USA 79:4108–4112
Piggot PJ, Hoch JA (1985) Revised genetic linkage map of Bacillus subtilis. Microbiol Rev 49:158–179
Scheer-Abramowitz J, Gryczan TJ, Dubnau D (1981) Origin and mode of replication of plasmids pE194 and pUB110. Plasmid 6:67–77
Scott JR (1984) Regulation of plasmid replication. Microbiol Rev 48:1–23
Stalker DM, Kolter R, Helinski DR (1982) Plasmid R6K DNA replication: I. Complete nucleotide sequence of an autonomously replicating segment. J Mol Biol 161:33–43
Vieira J, Messing J (1982) The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene 19:259–268
Yoshimura K, Ikenaka Y, Murai M, Tanabe M, Seki T, Oshima Y (1983a) Construction of a Bacillus subtilis cloning vehicle with heterologous DNA sequence. Gene 24:255–263
Yoshimura K, Yamamoto O, Seki T, Oshima Y (1983b) Distribution of heterogeneous and homologous plasmids in Bacillus spp. (corrected version). Appl Environ Microbiol 46:1268–1275
Author information
Authors and Affiliations
Additional information
Communicated by M. Takanami
Rights and permissions
About this article
Cite this article
Murai, M., Miyashita, H., Araki, H. et al. Molecular structure of the replication origin of a Bacillus amyloliquefaciens plasmid pFTB14. Mol Gen Genet 210, 92–100 (1987). https://doi.org/10.1007/BF00337763
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00337763