Abstract
The genetic architecture of the QT interval, defined as the period from onset of depolarisation to completion of repolarisation of the ventricular myocardium, is incompletely understood. Only a minor part of the QT interval variation in the general population has been linked to autosomal variant loci. Altered X chromosome dosage in humans, as seen in sex chromosome aneuploidies such as Turner syndrome (TS) and Klinefelter syndrome (KS), is associated with altered QTc interval (heart rate corrected QT), indicating that genes, located in the pseudoautosomal region 1 of the X and Y chromosomes may contribute to QT interval variation. We investigate the dosage effect of the pseudoautosomal gene SLC25A6, encoding the membrane ADP/ATP translocase 3 in the inner mitochondrial membrane, on QTc interval duration. To this end we used human participants and in vivo zebrafish models. Analyses in humans, based on 44 patients with KS, 44 patients with TS, 59 male and 22 females, revealed a significant negative correlation between SLC25A6 expression level and QTc interval duration. Similarly, downregulation of slc25a6 in zebrafish increased QTc interval duration with pharmacological inhibition of KATP channels restoring the systolic duration, whereas overexpression of SLC25A6 shortened QTc, which was normalized by pharmacological activation of KATP channels. Our study demonstrate an inverse relationship between SLC25A6 dosage and QTc interval indicating that SLC25A6 contributes to QT interval variation.
Similar content being viewed by others
Introduction
The electrocardiographic QT interval reflects depolarization and repolarization of the myocardium, and patients with an either prolonged or shortened QT interval are at risk of arrhythmias and sudden cardiac death1. The pathophysiology and genetic architecture of the QT interval is incompletely understood. Pathogenic variants in several autosomal genes encoding cardiac potassium channels have been identified as implicated in Mendelian inherited long QT syndrome and short QT syndrome2. However, only a minor part of the QT interval variation in the general population has been linked to common autosomal variant loci through genome-wide association studies3,4,5, leaving a major part of the QT interval variation genetically elusive.
In the early twentieth century, Henry Cuthbert Bazett demonstrated that the QT interval duration displayed sexual dimorphism, with women having a 6% longer average frequency corrected QT (QTc) interval than men6. This sexual dimorphism is believed to be driven by sex hormones7. Recently, extensive sex differences at the transcription level of both autosomal and sex chromosomal genes have been identified8,9 suggesting an impact of sex chromosome dosage in sexual dimorphism in health and diseases.
Sex chromosome aneuploidies (SCAs) constitute a valuable human model to advance knowledge of sex chromosome dosage in health and disease. Interestingly, studies of the QTc interval in patients with Turner syndrome (TS; 45,X) and Klinefelter syndrome (KS; 47,XXY) have revealed that the QTc interval is longer in patients with TS10,11,12, while patients with KS have a shortened QTc interval13,14. Notably, in a study of patients with KS, QTc interval was found to be significantly shorter in patients expressing a higher level of autosomal genes DOCK7 and GSTM2, the X chromosomal gene SMCA1, as well as pseudoautosomal genes CD99, SLC25A6, P2RY8, ZBED1, and GTPBP614. SLC25A6, Solute Carrier Family 25 Member 6, encodes the membrane ADP/ATP translocase 3, a membrane protein located in the inner mitochondrial membrane, which is involved in the exchange of intra-mitochondrial ATP for cytoplasmic ADP15, plays a key role in repiratory ATP synthesis, which may be critical for QTc duration. SLC25A6 is expressed in various levels in almost all human tissues, including cardiomyocytes16. Single cell transcriptomics of human heart tissue futher revealed that SLC25A6 is expressed in all cell types, including cardiomyocytes and smooth muscle cells (www.proteinatlas.org)17 (see Supplementary Fig. 1). Compared to female and male controls, TS patients show lower expression18,19, while patients with KS show higher expression of SLC25A614,18,19,20,21.
To determine a possible effect of altered dosage of SLC25A6 on the duration of the QT interval, we used both human participants including patients with TS and KS and zebrafish models. The zebrafish has been emphasized as a successful model to investigate the role of specific genes in human diseases and development22 and specifically in heart physiology23,24. Although the heart of zebrafish, like the heart from any common model organism, differs from the human heart, action potential in zebrafish cardiomyocytes shares the main characteristics of action potential of human ventricle myocytes25. The ionic current background of membrane potential changes in zebrafish cardiomyocytes differs slightly from humans, but the majority of ion channel orthologs important for human ventricular cardiomyocyte action potential are expressed in zebrafish heart. These include the sodium and calcium voltage-gated channels, the inward rectifying and ATP-dependent potassium channels. Therefore, cardiomyocyte action potential shape and duration result in heart rate-dependent QT-interval, which is highly similar between zebrafish and humans, making the zebrafish an attractive model organism for studying the effect of aberrant gene expression on cardiac physiology of relevance to humans23,24. The zebrafish genome contains a single highly conserved SLC25A6 orthologue, slc25a6, encoding a protein with 92% sequence identity to human SLC25A6. Thus, the zebrafish presents an attractive model to assess the consequences of altered SLC25A6 expression on cardiac rhythmicity in vivo.
In our case–control study, including TS, KS and female and male controls, we showed a significant negative correlation between the expression level of SLC25A6 and QTc interval duration. Similarly, in zebrafish, we observed that downregulation of slc25a6 resulted in a prolongation of the QTc interval whereas overexpression of SLC25A6 resulted in a shortening of the QTc interval. As a therapeutic intervention, we investigated the effect of pharmacological KATP channel inhibition in zebrafish with downregulated slc25a6 and KATP channel opening in zebrafish with overexpression of SLC25A6. Both interventions rescued the phenotypes. Collectively, our data shows that dosage of SLC25A6 impacts QTc interval and that these changes can bepharmacologically reversible.
Results
SLC25A6 expression levels correlate with sex-chromosome dosage in humans across tissue type of mesoderm origin
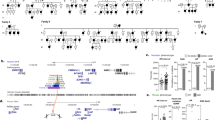
To determine the effect of altered dosage of SLC25A6 on QTc interval durations in humans, we first examined the relative expression level of SLC25A6 in peripheral blood samples from patients with TS, KS and female and male controls using quantitative PCR (see Supplementary Fig. 2, Supplementary Table S1). In agreement with recent published studies18,19, we found that TS was associated with a significant downregulation of SLC25A6, whereas KS was associated with a significant upregulation of SLC25A6 (Fig. 1A). We also found that SLC25A6 was significantly upregulated in male controls compared to female controls (Fig. 1A). This is in agreement with a previous study demonstrating that SLC25A6 exhibits a male-biased expression profile across human tissues, i.e., the expression profile mediated by an extension of the X inactivation process into the PAR region of the inactive X chromosome in individuals with more than one X chromosome26. To validate our data of relative expression levels, we assessed the expression level of SLC25A6 using RNA sequencing in a subpopulation of patients with TS and KS and in female and male controls. A significant correlation between our relative expression levels and absolute expression levels of SLC25A6 was seen (r2 = 0.60, p < 0.001) (Fig. 1B). To investigate if the expression pattern of SLC25A6 seen in blood is representative of other human tissues of mesodermal origin (heart, aorta, muscle), we analyzed the expression level of SLC25A6 using RNA sequencing in blood, muscle and aorta tissue from a subset of patients with TS and KS and female and male controls. The expression pattern of muscle tissue was comparable to that seen in blood with an upregulation in KS and downregulation in TS compared to controls (Supplementary Fig. 3) and analysis of the expression of SCL25A6 in aorta from patients with TS showed an expression level equal to the expression level in blood seen in patients with TS (see Supplementary Fig. 3).
SLC25A6 expression levels in humans correlate with QTc interval duration. (A) Box plot and table showing relative expression levels of SLC25A6 determined by qPCR in patients with Turner syndrome, Klinefelter syndrome and female and male controls. Values in table are medians and range. n (TS) = 54, n(KS) = 44, n(females) = 22, n(males) = 59. (B) Correlations between absolute expression values of SLC25A6 (counts) and relative expression of SLC25A6 in the subpopulation. n(TS) = 10, n(KS) = 6, n(females) = 7, n(males) = 6). (C) Box plot and table of QTc in patients with Turner syndrome, Klinefelter syndrome and female and male controls. Values in the table are median and range. n(TS) = 54, n(KS) = 44, n(females) = 22, n(males) = 59. (D) Correlation between QTc and relative expression of SLC25A6. n(TS) = 54, n(KS) = 44, n(females) = 22, n(males) = 59). Corrected QT interval (QTc) was calculated using Bazett´s equation (QTc = QT × √1/RR). Statistical analysis were performed by Mann Whitney U-test (median with range).TS, Turner syndrome. KS, Klinefelter syndrome. *p < 0.05, ***p < 0.001.
SLC25A6 expression levels in humans correlate with QTc interval duration
After having showed that SLC25A6 expression levels correlate with sex-chromosome dosage in humans across tissue type of mesodermal origin, we next analyzed QTc interval duration from previously recorded ECGs in our cohort of patients with TS, KS and female and male controls to be able to correlate SLC25A6 with QTc interval duration. We found that KS demonstrated the shortest QTc interval duration, followed by male controls and female controls, with TS showing the longest QTc interval duration (Fig. 1C). Correlation analysis of QTc interval duration and SLC25A6 expression demonstrated a significant negative correlation (r2 = − 0.33, p < 0.001) (Fig. 1D). However, the low r2 indicates that only a part of the variability seen in QTc interval can be explained by SLC25A6 expression level.
SLC25A6 expression level impacts systolic duration and QTc interval duration in zebrafish
To gain further evidence of an impact of SLC25A6 expression level on QTc interval, we confirmed cardiac slc25a6 expression by RT-PCR on a pool of isolated 3 days post fertilization (dpf) zebrafish hearts (see Supplementary Fig. 4) and recorded the effect of SLC25A6 overexpression and slc25a6 downregulation on systolic contraction duration in 3 days post fertilization (dpf) zebrafish larvae using high-speed video recordings. The physiological condition of KS carrying three copies of SLC25A6 was modeled in zebrafish embryos by overexpression of wildtype SLC25A6 by mRNA microinjection into newly fertilized zebrafish embryos. RT-PCR analysis indicated similar in vivo stability of human wildtype SLC25A6 mRNA and full length control SLC25A6 mRNA containing two premature stop codons (see Supplementary Fig. 5). The physiological condition of TS carrying only one copy of SLC25A6 was modeled in zebrafish embryos by slc25a6 knockdown titrated to approximately 50% downregulation of the wildtype endogenous slc25a6 mRNA level (see Supplementary Fig. 6). Neither SLC25A6 overexpression nor slc25a6 downregulation overtly affected zebrafish embryo morphology (see Supplementary Fig. 7).
We observed that in zebrafish with overexpression of SLC25A6, a significant shortening of systolic duration occurred compared to controls (Fig. 2A). In addition, slc25a6 downregulation resulted in a significant prolongation of systolic duration compared to controls (Fig. 2C). No effect on heart rate was seen (Fig. 2B,D,F,H). The specificity of the phenotype of slc25a6 downregulated embryos was substantiated by phenotypic rescue resulting from the co-injection of wildtype SLC25A6 mRNA, but not control mRNA (Fig. 2C). Importantly, ECG analysis of 3 days post fertilization zebrafish larvae revealed a significantly shortened and prolonged QTc interval duration by SLC25A6 overexpression and slc25a6 downregulation, respectively (Fig. 2E,G), demonstrating that the observed changes in systolic duration correspond to altered QTc interval duration. The averaged ECG recordings and representative ECG traces are shown in Supplementary Fig. 9. Collectively, these findings show that that the level of SLC25A6 expression impact QTc interval duration in vivo, thereby adding futher evidence of an impact of SLC25A6 expression level on QTc interval.
SLC25A6 expression level impacts systolic duration and QTc interval duration in zebrafish. (A) Systolic interval normalized to heart period determined from high-speed video recordings of the heart ventricle show reduced systolic duration resulting from SLC25A6 overexpression (SLC25A6 mRNA) compared to controls (Control mRNA). (B) SLC25A6 overexpression did not affect heart rate. Data in (A,B) were compiled from three independent experiments, each encompassing both groups (n = 15). (C) Systolic interval normalized to heart period determined from high-speed video recordings of the heart ventricle show increased systolic duration resulting from approximately 50% knockdown of endogenous slc25a6 mRNA (slc25a6 KD) compared to controls (Control KD). The phenotype was rescued by co-injection of the slc25a6 targeted morpholino with wildtype SLC25A6 mRNA (slc25a6 KD + SLC25A6 mRNA) but not control mRNA (slc25a6 KD + control mRNA). (D) Knockdown of endogenous slc25a6 mRNA did not affect heart rate. Data in (C,D) were compiled from three independent experiments, each encompassing all four groups. n(control KD) = 13, n(slc25a6 KD) = 10, n(slc25a6 KD + control mRNA) = 13, n(slc25a6 KD + SLC25A6 mRNA) = 12. (E) SLC25A6 overexpression (SLC25A6 mRNA) resulted in a decreased QTc interval compared to controls (Control mRNA). (G) 50% downregulation of endogenous slc25a6 mRNA (slc25a6 KD) resulted in an increased QTc interval compared to controls (Control KD). (F,H) SLC25A6 overexpression (SLC25A6 mRNA) and downregulation of endogenous slc25a6 (slc25a6 KD) did not affect heart rate. n = 15 (Control mRNA), n = 15 (SLC25A6 mRNA), n = 14 (Control KD), n = 1 (slc25a6 KD). Corrected QT interval (QTc) was calculated using Bazett’s equation (QTc = QT × √1/RR). Statistical analyses were performed by one-way ANOVA with Tukey’s post-test. *p < 0.05, ***p < 0.001.
Pharmacological rescue of SLC25A6 overexpression and slc25a6 knockdown phenotypes
As SLC25A6 doses can modify intracellular ADP/ATP ratio through its ADP/ATP antiporter activity, we hypothesized an involvement of the KATP channel. To test this hypothesis, we treated slc25a6 knockdown zebrafish with the KATP channel antagonist glybenclamide and quantified systolic duration. We observed that glybenclamide treatment resulted in a normalization of the systolic duration (Fig. 3A). Heart rate was unaffected by glybenclamide treatment (Fig. 3B) and no pharmacological effect of glybenclamide on controls was observed (p = 0.50) (Fig. 3A), in accordance with the majority of KATP channels consitutevely closed under resting conditions27. Treatment with KATP channel agonist pinacidil normalized the shortened systolic interval duration induced by SLC25A6 overexpression with no pharmacological effect on controls (Fig. 3C). Heart rate was unaffected by pinacidil treatment (Fig. 3D). Collectively, these results show, that prolonged or shortened systolic interval caused by altered SLC25A6 expression can be modulated pharmacologically.
Pharmacological rescue of SLC25A6 overexpression and slc25a6 knockdown phenotypes. (A) Systolic duration normalized to heart period determined from high-speed video recordings of the heart ventricle. Systolic interval prolongation induced by slc25a6 knockdown (slc25a6 KD) is normalized by glybenclamide treatment (slc25a6 KD + glybenclamide). (B) Neither slc25a6 knockdown (slc25a6 KD) nor glybenclamide treatment (control KD + glybenclamide and slc25a6 KD + glybenclamide) affects heart rate. Data in (A,B) were compiled from four independent experiments, each encompassing all four groups. n(Control KD) = 28, n(Control KD + glybenclamide) = 37, n(slc25a6 KD) = 26, n(slc25a6 KD + glybenclamide) = 25. (C) Systolic interval normalized to heart period determined from high-speed video recordings of the heart ventricle. Systolic interval shortening induced by SLC25A6 overexpression (SLC25A6 mRNA) is diminished by pinacidil treatment (SLC25A6 mRNA + pinacidil). (D) Neither SLC25A6 overexpression (SLC25a6 mRNA) nor pinacidil treatment (Control mRNA + pinacidil and SLC25A6 mRNA + pinacidil) affects heart rate. Data in (C,D) were compiled from three independent experiments, each encompassing all four groups. n(Control mRNA) = 18, n(Control mRNA + pinacidil) = 16, n(SLC25A6 mRNA) = 15, n(SLC25A6 mRNA + pinacidil) = 17. Statistical analyses were performed by one-way ANOVA with Tukey´s post-test. *p < 0.05, **p < 0.01, ****p < 0.0001.
Discussion
This study finds evidence of the first pseudoautosomal gene, SLC25A6, contributing to QTc interval duration. We showed a significant negative correlation between the expression of SLC25A6 and QTc interval in humans. Our zebrafish model supported these findings and further revealed that pharmacological intervention was possible. Our data not only provide evidence that SLC25A6 contributes to QT interval variation in the general population, but also that SLC25A6 is involved in the cardiac phenotypes with altered QTc interval seen in TS and KS.
The significant correlation found between the expression level of SLC25A6 and QTc interval across humans adds SLC25A6 to the list of genes contributing to the variation in QT interval seen in the human population. Previously, genome-wide association (GWAS) studies have contributed to the identification of common autosomal variants, which explain up to 8–10% of QT interval variation in the general population. However, one of the limitations of previously performed GWAS studies is the inclusion of only autosomal variants, an approach overlooking the possibility that pseudoautosomal genes and X chromosomal genes could contribute to QT interval variation. Our study highlights the role of the pseudoautosomal gene SLC25A6 in explaining QT interval variation; however other pseudoautosomal and X chromosomal genes may also be involved.
The rescuing effect of the KATP channel agonist pinacidil and the KATP channel antagonist glybenclamide on systolic interval duration in our zebrafish model indicated that expression levels of SLC25A6 might indirectly affect the function of the KATP channel in the cardiomyocytes through alterations of ATP/ADP levels. The role of KATP channels in cardiac electrical activity is rather complex and depends on their localization28,29. The KATP channels in ventricle cardiomyocytes are generally considered cardioprotective metabolic sensors that activate upon acute ischemic conditions and shorten the action potential duration and thus QT interval. These changes normally occur because of severe changes in intracellular ATP upon metabolic impairment. In contrast, the KATP channels of the cardiac conduction system have a different molecular composition that makes them more readily activated by ADP and less sensitive to inhibitory action of ATP, i.e., sensitive to minor changes in ADP and ATP30. Activation of these KATP channels may slow the conduction velocity and thus, prolong the systolic duration and QT interval31 as seen under chronic ischemic conditions32. Moreover, gain-of-function mutations in the KATP channel were suggested to prolong repolarization in some (epicardial) cardiomyocytes33, as well as increasing potassium transmembrane gradient, while reduction of the gradient shortened action potential34.
In recent years, our knowledge of the biology of ADP/ATP translocases has expanded. In addition to their involvement in ATP/ADP exchange, evidence also suggests that ADP/ATP translocases are targets of mitochondrial uncouplers35 and involved in mitochondrial H+ leak36, mitochondrial membrane potential maintenance37, and potentiation of the mitochondrial permeability transition pore38. It is therefore possible, that the alterations seen in QTc interval with altered SLC25A6 expression, are due to a more general impact of SLC25A6 on the mitochondrial function, and not an isolated effect on the KATP channel. Future studies are needed to demonstrate the mechanistic contribution of the ADP/ATP translocase to electrical function of cardiomyocytes.
Further studies will be needed to decipher the molecular mechanisms and physiological basis of the SLC25A6 dosage and expression level-dependent regulation of QT interval variation.
In conclusion, our study highlights that the dosage and expression level of the pseudoautosomal gene SLC25A6 contribute to QTc interval variation.
Methods
Human subjects
The human subjects were recruited as previously described11,13. Males with verified KS (n = 44) were recruited through endocrine and clinical genetic hospital outpatient clinics as well as fertility clinics in Denmark, whereas women with verified TS (n = 54) were recruited through the Danish National Society of Tuner Syndrome Contact Group and an endocrine outpatient clinic in Denmark. Healthy, age-matched men (n = 59) and women (n = 22) were recruited through advertisement to serve as controls. The subpopulation included males with verified KS (n = 6), women with verified TS (n = 10), healthy, age-matched men (n = 6) and women (n = 7). The validation cohort included males with verified KS (n = 16), women with verified TS (n = 10), healthy men (n = 16) and women (n = 5), all included in the main population. Participants were recruited as described above.
Relative quantification of SLC25A6 expression in peripheral blood from human subjects
Two and a half ml of peripheral blood were collected in PaxGene blood RNA tubes (total volume 10 ml) (BD, Franklin Lakes, NJ, USA) followed by extraction of total RNA using PaxGene Blood RNA Kit (Qiagen, Hilden, Germany) according to the manufacturer’s protocol including DNAase I treatment to remove residual genomic DNA. The RNA was eluted in 40 µl of nuclease-free water, quantified using Dropsense96 (PerkinElmer, Waltham, MA, USA) and stored at − 80 °C. Subsequently, 500 ng of RNA was converted to cDNA using SuperScript II reverse transcriptase (ThermoFisher, Waltham, MA, USA) according to the manufacturers recommendations and a combination of Oligo(dT)15 (250 ng) (5ʹ-TTT TTT TTT TTT TTT VN-3ʹ, V = A, G or C, and N = A, T, G or C (SigmaAldrich, St Louis, MO, USA)) and random hexamer primers (Invitrogen, Carlsbad, CA, USA) (125 ng; reaction volume 20 µl). The expression of SLC25A6 and Ubiquitin C (UBC) (endogenous reference gene) were measured using quantitative PCR (qPCR). The qPCR reactions were performed in triplicates on a ViiA 7 Real-Time PCR instrument (Applied Biosystems, Thermofisher, Waltham, MA, USA) according to the manufacturer’s instructions and using the following TaqMan gene expression assays (ThermoFisher): Hs00745067_s1 (SLC25A6) and Hs00824723_m1 (UBC). 2 μl of each cDNA (diluted 50-fold) was used as a template in the qPCR reactions together with 8 μl Gene Expression Master Mix (reaction volume 10 µl). Representative qPCR curves are illustrated in Supplementary Fig. 2). Raw CT values (Supplementary Table S1) were calculated with automatic baseline and threshold settings, and the relative expression levels of SLC25A6 were determined using the comparative CT method (2−ΔΔCT method), i.e., normalization to the reference gene (UBC) and relative to an internal calibrator sample using the QuantStudio Real-Time PCR Software v1.3 (Thermofisher).
Whole blood SLC25A6 expression in the subpopulation
SLC25A6 expression was assessed in a separate cohort based on RNA sequencing of whole blood from females with verified TS and healthy women (Trolle et al.39, data accessible at the European Genome-Phenome Archive (EGA), EGAS00001002190), and males with verified KS and healthy men (Skakkebæk et al.21, data accessible at the EGA, EGAS00001002797). Paired de-multiplexed fastq files were subjected to initial quality control using FastQC (Babraham Bioinformatics). The adaptor removal, in addition to trimming of low-quality ends, was then conducted using Trim Galore with default settings (Babraham Bioinformatics), followed by mapping to the human genome (hg38) using HISAT240. The overlaps between genomic annotations (Gencode v 37) and the read alignments were quantified by feature Counts to produce gene counts for each sample41. The read counts were then normalized by the R Bioconductor package DESeq242, and the estimated gene expression of SLC25A6 for each sample was used for correlation analysis.
Whole blood and muscle tissue expression of SLC25A6 in the validation cohort
SLC25A6 expression was assessed in a separate cohort based on RNA sequencing of whole blood and muscle from females with verified TS, males with verified KS and healthy women and men43. The RNA-seq libraries were multiplexed paired-end sequenced on a Illumina NovaSeq 6000 (100 bp). Paired de-multiplexed fastq files were subjected to initial quality control using FastQC (Babraham Bioinformatics). Adaptor removal and trimming of low-quality ends were then conducted using Trim Galore with default settings (Babraham Bioinformatics). Gene expression was quantified by quasi-mapping using Salmon44. A decoy-aware transcriptome index was built based on the hg38 transcriptome and selective alignment was run using the fastq pairs as input. Transcript abundancies were summarized to gene-level using the R package Tximeta45. The gene counts were then normalized by the R Bioconductor package DESeq242, and the estimated gene expression of SLC25A6 for each sample was used for correlation analysis.
Aortic expression of SLC25A6
Aortic biopsies were collected from three patients with TS undergoing planned elective thoracic surgery due to aortic dilatation. The biopsies were immediately cleaned and snap-frozen in liquid nitrogen and stored at − 80 °C. The frozen biopsies were dissociated using a CovarisSP02 and RNA was extracted by the AllPrep DNA/RNA/PROTEIN Mini kit 8004 (Qiagen). Synthesis of directional RNA-seq libraries were conducted as described above using the KAPA RNA HyperPrep with RiboErase kit (HMR) (Roche).
Electrocardiogram and QTc measurements in human subjects
Twelve lead electrocardiograms (ECG) from females with verified TS and healthy women, and, males with verified KS and healthy men were obtained as described previously11,13. ECGs were recorded at 25 mm/s with an amplitude of 10 mm/mV (Personal 120/210 machine (Esaote Biomedica, Cambridge, United Kingdom) and scanned to a digital file. Cardio Calipers 3.3. (Iconico, http://www.iconico.com) were used to measure QT interval and RR interval. The corrected QT interval (QTc) was calculated using Bazett´s equation (QTc = QT × √1/RR)6.
Zebrafish
Adult AB wildtype zebrafish originating from the European Zebrafish Resource Center (Karlsruhe, Germany) were maintained on commercial recirculating zebrafish housing systems (Tecniplast, Buguggiate, Italy) in reverse osmosis water conditioned to 0.7mS, pH 7.2, 28 °C, on a 14 h light/10 h darkness cycle and fed four times daily. Embryos were obtained by natural crosses and reared in E3 medium (5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl2, 0.33 mM MgSO4, 10–5% methylene blue, 2 mM HEPES adjusted to pH 7.4 with NaHCO3) at 28 °C.
Overexpression of SLC25A6 and knockdown of slc25a6 in zebrafish
To evaluate embryonic cardiac slc25a6 expression, hearts from 12.3 days post fertilization (dpf) zebrafish embryos were isolated by manual microdissection and pooled in RNAlater (Sigma). RNA was isolated using Qiagen RNeasy Plus Micro Kit. RT-PCR was performed using Qiagen Onestep RT-PCR Kit and primers pairs 5ʹ-CACGAGACCACCTTCAACT-3ʹ/5ʹ-ATCCAGACGGAGTATTTGC-3ʹ (β-actin) and 5ʹ-TGCAGCCTTTTCTTTTGCGA-3ʹ/5ʹ-AATGGGTAGGAGACCACGCC-3ʹ (slc25a6) followed by a nested PCR reaction using primer pair 5ʹ-CAGATGGCTTACGAGGGCTG-3ʹ/5ʹ-TGTGGGTGTTCTTTGGATCGG-3ʹ (slc25a6). PCR product identity were verified by Sanger sequencing. Primer pairs 5ʹ AATTAATACGACTCACTATAGAACATGACGGAACAGGCCATCTC/CACACTGGACTAGTGGATCCTTAG-3ʹ and 5ʹ AATTAATACGACTCACTATAGAACATGACGTAATAGGCCATCTC-3ʹ/5ʹ-CACACTGGACTAGTGGATCCTTAG-3ʹ were used to generate T7-promoter-containing Kozak-optimized templates for the synthesis of mRNA encoding wildtype human SLC25A6 and a full length control SLC25A6 transcript containing two premature stop codons (SLC25A6 E3stop,Q4stop), using Genscript clone OHu19558 as a template. mRNAs were synthesized using mMessage mMachine T7 ULTRA kit (Ambion) according to suppliers’ recommendations and microinjected at 400 pg per embryo into freshly fertilized zebrafish embryos no later than the two-cell stage. The stability of injected transcripts 3 days post fertilization was assessed by RT PCR using primers: 5ʹ-CAAGAACACGCACATCGTGG-3ʹ/5ʹ-GTACAGGACCAGCACGAAGG-3ʹ and β-actin mRNA levels were assessed using the primer pair 5ʹ-CACGAGACCACCTTCAACT-3ʹ/5ʹ-ATCCAGACGGAGTATTTGC-3ʹ. PCR product identities were verified by Sanger sequencing. slc25a6 knockdown was performed by microinjection of slc25a6 exon2-intron2 targeted morpholino (TCACAAACCTATTACCTTTAGCAGT) into newly fertilized zebrafish embryos no later than the two-cell stage. The morpholino was titrated to obtain an approximately 50% reduction of endogenous slc25a6 mRNA levels at 3 dpf. Standard control morpholino (CCTCTTACCTCAGTTACAATTTATA, Gene Tools, LLC) was injected in equimolar amount as a negative control. Morpholino efficiency was assessed by reverse transcription PCR (RT-PCR) from cDNA synthesized from total RNA extracted from groups of 10 microinjected zebrafish larvae three days post fertilization using the primer pair 5ʹ-GCGCCGATTGAGAGAGTCAA-3ʹ/5ʹ-AATGGGTAGGAGACCACGCC-3ʹ and PCR product identities were verified by Sanger sequencing. Band intensities were determined using ImageJ46.
Imaging and measurements of cardiac function and ECG in zebrafish
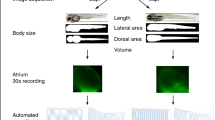
Whole-mount images were done on 3 days post fertilization zebrafish larvae anaesthetized with 150 ng/ml ethyl 3-aminobenzoate methanesulfonate (150 ng/ml; MS-222; Sigma-Aldrich) in E3 medium and staged in 3% hydroxypropyl methylcellulose (average Mn ~ 86,000, Sigma Aldrich) in E3 medium. High-speed video recordings of cardiac function of zebrafish larvae were performed (Supplementary Fig. 8) and analyzed as previously described47.
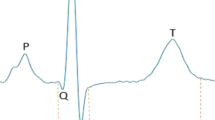
ECG recordings in 3 days post fertilization zebrafish larvae were performed as described previously47. Zebrafish larvae were anaesthetized using 150 ng/ml MS-222 (Sigma) in E3 medium at room temperature (22–24 °C). A borosilicate glass micropipette (PG15OT-7.5; Harvard Apparatus) filled with 3 mM KCl with tip resistances of approximately 10 MΩ was positioned on larvae skin surface (no penetration). Data acquisition was done at 10 kHz using an Axopatch 200B amplifier (Axon Instruments, Inc.) in a current-clamp configuration with the software package Clampex 7 (Axon Instruments Inc.). Data analysis was automated in ECG analysis module of LabChart 8 software (ADInstruments) (Supplementary Fig. 9). QT interval and RR interval were automatically calculated for each larva and the corrected QT interval (QTc) was assessed using Bazett’s equation (QTc = QT/√RR, where RR is the interval between two consequent R waves in ECG)6.
Pharmacological interventions with glybenclamide and pinacidil in zebrafish
For pharmacological rescue experiments, 3 days post fertilization zebrafish larvae were treated with 50 µM pinacidil (Sigma-Aldrich P154), 50 µM glybenclamide (Sigma-Aldrich G0639) in 0.1% DMSO or vehicle control combined with 150 ng/ml MS-222 (Sigma) anesthesia in E3 medium 15–30 min prior to high-speed recordings.
Statistics
Statistical analyses were conducted using SPSS 21.0 (IBM Corp., Armonk, NY, USA) and GraphPad Prism 7 (GraphPad Software, San Diego, CA). Comparisons of continuous variables were performed using Student’s independent t test (mean ± SD) or one-way ANOVA with Tukey´s post-test for normally distributed variables, and Mann Whitney U-test (median with range) was used for non-parametric variables.
Study approval
The study was approved by The Danish Data Protection Agency and the local and central ethics committee (Region Midtjylland, Denmark number M-20080238 and M-20010248, Central Denmark Regional Committee on Health Research Ethics number 1-10-72-131-15) and registered at ClinicalTrials.gov (NCT00624949, NCT00999310, NCT02526628). All participants provided informed consent. All methods were performed in accordance with the relevant guidelines and regulations. All experimental zebrafish procedures were performed in agreement with Danish legislation and guidelines from the European Convention for the Protection of Vertebrate Animals used for Experimental and other Scientific Purposes. The study was reported in accordance with ARRIVE guidelines.
Data availability
The data supporting the findings of this study are available from the corresponding author upon reasonable request. The RNA sequencing datasets used in this study are available at the European Genome-Phenome Archive (EGA) (EGAS00001002190, EGAS00001002797, EGAS00001006404).
References
Scrocco, C., Bezzina, C. R., Ackerman, M. J. & Behr, E. R. Genetics and genomics of arrhythmic risk: Current and future strategies to prevent sudden cardiac death. Nat. Rev. Cardiol. 18, 774–784 (2021).
Giudicessi, J. R. & Ackerman, M. J. Potassium-channel mutations and cardiac arrhythmias-diagnosis and therapy. Nat. Rev. Cardiol. 9, 319–332 (2012).
Arking, D. E. et al. Genetic association study of QT interval highlights role for calcium signaling pathways in myocardial repolarization. Nat. Genet. 46, 826–836 (2014).
Newton-Cheh, C. et al. Common variants at ten loci influence QT interval duration in the QTGEN study. Nat. Genet. 41, 399–406 (2009).
Noseworthy, P. A. et al. Common genetic variants, QT interval, and sudden cardiac death in a Finnish population-based study. Circ. Cardiovasc. Genet. 4, 305–311 (2011).
Bazett, H. C. An analysis of the time-relations of electrocardiograms. Heart 7, 18 (1920).
Offerhaus, J. A., Bezzina, C. R. & Wilde, A. A. M. Epidemiology of inherited arrhythmias. Nat. Rev. Cardiol. 17, 205–215 (2020).
Gershoni, M. & Pietrokovski, S. The landscape of sex-differential transcriptome and its consequent selection in human adults. BMC Biol. 15, 7. https://doi.org/10.1186/s12915-017-0352-z (2017).
Oliva, M. et al. The impact of sex on gene expression across human tissues. Science 369, 3066 (2020).
Bondy, C. A. et al. Prolongation of the cardiac QTc interval in Turner syndrome. Medicine 85, 75–81 (2006).
Trolle, C. et al. Long QT interval in Turner syndrome—A high prevalence of LQTS gene mutations. PLoS ONE 8, e69614. https://doi.org/10.1371/journal.pone.0069614 (2013).
Sozen, A. B. et al. Atrial and ventricular arryhthmogenic potential in Turner syndrome. Pace 31, 1140–1145 (2008).
Jorgensen, I. N. et al. Short QTc interval in males with Klinefelter syndrome-influence of CAG repeat length, body composition, and testosterone replacement therapy. Pacing Clin. Electrophysiol. 38, 472–482 (2015).
Zitzmann, M. et al. Gene expression patterns in relation to the clinical phenotype in Klinefelter syndrome. J Clin. Endocrinol. Metab. 100, E518–E523 (2015).
Slim, R. et al. A human pseudoautosomal gene encodes the ANT3 ADP/ATP translocase and escapes X-inactivation. Genomics 16, 26–33 (1993).
Stepien, G., Torroni, A., Chung, A. B., Hodge, J. A. & Wallace, D. C. Differential expression of adenine nucleotide translocator isoforms in mammalian tissues and during muscle cell differentiation. J. Biol. Chem. 267, 14592–14597 (1992).
Karlsson, M. et al. A single-cell type transcriptomics map of human tissues. Sci. Adv. 7, 2169. https://doi.org/10.1126/sciadv.abh2169 (2021).
Zhang, X. et al. Integrated functional genomic analyses of Klinefelter and Turner syndromes reveal global network effects of altered X chromosome dosage. Proc. Natl. Acad. Sci. U.S.A. 117, 4864–4873 (2020).
Manotas, M. C. et al. Identification of common differentially expressed genes in Turner (45, X) and Klinefelter (47, XXY) syndromes using bioinformatics analysis. Mol. Genet. Genom. Med. 8, e1503. https://doi.org/10.1002/mgg3.1503 (2020).
Belling, K. et al. Klinefelter syndrome comorbidities linked to increased X chromosome gene dosage and altered protein interactome activity. Hum. Mol. Genet. 26, 1219–1229 (2017).
Skakkebaek, A. et al. DNA hypermethylation and differential gene expression associated with Klinefelter syndrome. Sci. Rep. 8, 13740. https://doi.org/10.1038/s41598-018-31780-0 (2018).
Howe, K. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 496, 498–503 (2013).
Nemtsas, P., Wettwer, E., Christ, T., Weidinger, G. & Ravens, U. Adult zebrafish heart as a model for human heart? An electrophysiological study. J. Mol. Cell. Cardiol. 48, 161–171 (2010).
Echeazarra, L., Hortigon-Vinagre, M. P., Casis, O. & Gallego, M. Adult and developing zebrafish as suitable models for cardiac electrophysiology and pathology in research and industry. Front. Physiol. 11, 607860. https://doi.org/10.3389/fphys.2020.607860 (2020).
Vornanen, M. & Hassinen, M. Zebrafish heart as a model for human cardiac electrophysiology. Channels 10, 101–110 (2016).
Tukiainen, T. et al. Landscape of X chromosome inactivation across human tissues. Nature 550, 244–248 (2017).
Kantor, P. F., Coetzee, W. A., Carmeliet, E. E., Dennis, S. C. & Opie, L. H. Reduction of ischemic K+ loss and arrhythmias in rat hearts. Effect of glibenclamide, a sulfonylurea. Circ. Res. 66, 478–485 (1990).
Nichols, C. G. Adenosine triphosphate-sensitive potassium currents in heart disease and cardioprotection. Card. Electrophysiol. Clin. 8, 323–335 (2016).
Foster, M. N. & Coetzee, W. A. KATP Channels in the cardiovascular system. Physiol. Rev. 96, 177–252 (2016).
Bao, L. et al. Unique properties of the ATP-sensitive K(+) channel in the mouse ventricular cardiac conduction system. Circ. Arrhythm. Electrophysiol. 4, 926–935 (2011).
Tse, G., Chan, Y. W., Keung, W. & Yan, B. P. Electrophysiological mechanisms of long and short QT syndromes. Int. J. Cardiol. Heart Vasc. 14, 8–13 (2017).
Beinart, R. et al. The QT interval is associated with incident cardiovascular events: The MESA study. J. Am. Coll. Cardiol. 64, 2111–2119 (2014).
Kusano, K. F. Brugada syndrome: Recent understanding of pathophysiological mechanism and treatment. J. Arrhythmia 29, 77–82 (2013).
Yan, G. X. & Antzelevitch, C. Cellular basis for the normal T wave and the electrocardiographic manifestations of the long-QT syndrome. Circulation 98, 1928–1936 (1998).
Bertholet, A. M. et al. Mitochondrial uncouplers induce proton leak by activating AAC and UCP1. Nature 606, 180–187 (2022).
Bertholet, A. M. et al. H(+) transport is an integral function of the mitochondrial ADP/ATP carrier. Nature 571, 515–520 (2019).
Ouyang, Y. et al. Phosphate starvation signaling increases mitochondrial membrane potential through respiration-independent mechanisms. BioRxiv 11, e1004931 (2022).
Karch, J. et al. Inhibition of mitochondrial permeability transition by deletion of the ANT family and CypD. Sci. Adv. 5, 4597. https://doi.org/10.1126/sciadv.aaw4597 (2019).
Trolle, C. et al. Widespread DNA hypomethylation and differential gene expression in Turner syndrome. Sci. Rep. 6, 34220. https://doi.org/10.1038/srep34220 (2016).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Johannsen, E. B. et al. Sex chromosome aneuploidies give rise to changes in the circular RNA profile: A circular transcriptome-wide study of Turner and Klinefelter syndrome across different tissues. Front. Genet. 13, 928874. https://doi.org/10.3389/fgene.2022.928874 (2022).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I. et al. Tximeta: Reference sequence checksums for provenance identification in RNA-seq. PLoS Comput. Biol. 16, e1007664. https://doi.org/10.1371/journal.pcbi.1007664 (2020).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Thorsen, K. et al. Loss-of-activity-mutation in the cardiac chloride-bicarbonate exchanger AE3 causes short QT syndrome. Nat. Commun. https://doi.org/10.1038/s41467-017-01630-0 (2017).
Acknowledgements
The authors thank C. Trolle, M.G.B. Pedersen, K.M. Lauritsen for help with fat and muscle tissue biopsies. They thank Lone Kvist and Pamela Celis for their technical assistance.
Funding
A.S. was supported by the Independent Research Fund Denmark (0134-00130B), C.H.G was supported by Novo Nordisk Foundation (NNF15OC0016474, NNF20OC0060610), “Fonden til lægevidenskabens fremme”, the Familien Hede Nielsen foundation and the Independent Research Fund Denmark (0134-00406A).
Author information
Authors and Affiliations
Contributions
A.S., K.K.S., V.V.M., C.O. and C.G. designed the experiments; A.S., K.K.S., V.V.M., L.L.C. performed the experiments; A.S., K.K.S., V.V.M., L.L.C., and J.J. analyzed the data; A.S., K.K.S., V.V.M., L.L.C., J.J., C.C., N.H.A., C.O., C.H.G. wrote the paper. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Skakkebæk, A., Kjær-Sørensen, K., Matchkov, V.V. et al. Dosage of the pseudoautosomal gene SLC25A6 is implicated in QTc interval duration. Sci Rep 13, 12089 (2023). https://doi.org/10.1038/s41598-023-38867-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-38867-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.