Abstract
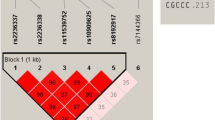
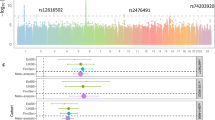
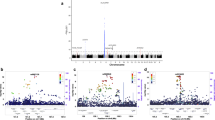
We conducted a genome-wide association study of generalized vitiligo in the Chinese Han population by genotyping 1,117 cases and 1,429 controls. The 34 most promising SNPs were carried forward for replication in samples from individuals of the Chinese Han (5,910 cases and 9,916 controls) and Chinese Uygur (713 cases and 824 controls) populations. We identified two independent association signals within the major histocompatibility complex (MHC) region (rs11966200, Pcombined = 1.48 × 10−48, OR = 1.90; rs9468925, Pcombined = 2.21 × 10−33, OR = 0.74). Further analyses suggested that the strong association at rs11966200 might reflect the reported association of the HLA-A*3001, HLA-B*1302, HLA-C*0602 and HLA-DRB1*0701 alleles and that the association at rs9468925 might represent a previously unknown HLA susceptibility allele. We also identified one previously undescribed risk locus at 6q27 (rs2236313, Pcombined = 9.72 × 10−17, OR = 1.20), which contains three genes: RNASET2, FGFR1OP and CCR6. Our study provides new insights into the genetic basis of vitiligo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hann, S.K. & Nordlund, J.J. Vitiligo (Blackwell Science, Oxford, UK, 2000).
Das, S.K., Majumder, P.P., Chakraborty, R., Majumdar, T.K. & Haldar, B. Studies on vitiligo. I. Epidemiological profile in Calcutta, India. Genet. Epidemiol. 2, 71–78 (1985).
Birlea, S.A., Fain, P.R. & Spritz, R.A. A Romanian population isolate with high frequency of vitiligo and associated autoimmune diseases. Arch. Dermatol. 144, 310–316 (2008).
Taïeb, A. & Picardo, M. Clinical practice. Vitiligo. N. Engl. J. Med. 360, 160–169 (2009).
Porter, J., Beuf, A.H., Nordlund, J.J. & Lerner, A.B. Psychological reaction to chronic skin disorders: a study of patients with vitiligo. Gen. Hosp. Psychiatry 1, 73–77 (1979).
Alkhateeb, A., Fain, P.R., Thody, A., Bennett, D.C. & Spritz, R.A. Epidemiology of vitiligo and associated autoimmune diseases in Caucasian probands and their families. Pigment Cell Res. 16, 208–214 (2003).
Dell'anna, M.L. & Picardo, M. A review and a new hypothesis for non-immunological pathogenetic mechanisms in vitiligo. Pigment Cell Res. 19, 406–411 (2006).
Spritz, R.A. The genetics of generalized vitiligo. Curr. Dir. Autoimmun. 10, 244–257 (2008).
Zhang, X.J. et al. Characteristics of genetic epidemiology and genetic models for vitiligo. J. Am. Acad. Dermatol. 51, 383–390 (2004).
Nath, S.K. et al. Evidence for a susceptibility gene, SLEV1, on chromosome 17p13 in families with vitiligo-related systemic lupus erythematosus. Am. J. Hum. Genet. 69, 1401–1406 (2001).
Alkhateeb, A. et al. Mapping of an autoimmunity susceptibility locus (AIS1) to chromosome 1p31.3–p32.2. Hum. Mol. Genet. 11, 661–667 (2002).
Spritz, R.A., Gowan, K., Bennett, D.C. & Fain, P.R. Novel vitiligo susceptibility loci on chromosomes 7 (AIS2) and 8 (AIS3), confirmation of SLEV1 on chromosome 17, and their roles in an autoimmune diathesis. Am. J. Hum. Genet. 74, 188–191 (2004).
Chen, J.J. et al. A novel linkage to generalized vitiligo on 4q13–q21 identified in a genomewide linkage analysis of Chinese families. Am. J. Hum. Genet. 76, 1057–1065 (2005).
Jin, Y. et al. NALP1 in vitiligo-associated multiple autoimmune disease. N. Engl. J. Med. 356, 1216–1225 (2007).
Ren, Y. et al. Genetic variation of promoter sequence modulates XBP1 expression and genetic risk for vitiligo. PLoS Genet. 5, e1000523 (2009).
Jin, Y., Birlea, S.A., Fain, P.R. & Spritz, R.A. Genetic variations in NALP1 are associated with generalized vitiligo in a Romanian population. J. Invest. Dermatol. 127, 2558–2562 (2007).
Zamani, M. et al. Linkage and association of HLA class II genes with vitiligo in a Dutch population. Br. J. Dermatol. 145, 90–94 (2001).
Arcos-Burgos, M. et al. Vitiligo: complex segregation and linkage disequilibrium analyses with respect to microsatellite loci spanning the HLA. Hum. Genet. 110, 334–342 (2002).
Jin, Y. et al. Variant of TYR and autoimmunity susceptibility loci in generalized vitiligo. N. Engl. J. Med. 362, 1686–1697 (2010).
Liu, J.B. et al. Association of vitiligo with HLA-A2: a meta-analysis. J. Eur. Acad. Dermatol. Venereol. 21, 205–213 (2007).
Zhang, X.J. et al. Association of HLA class I alleles with vitiligo in Chinese Hans. J. Dermatol. Sci. 35, 165–168 (2004).
Taştan, H.B., Akar, A., Orkunoglu, F.E., Arca, E. & Inal, A. Association of HLA class I antigens and HLA class II alleles with vitiligo in a Turkish population. Pigment Cell Res. 17, 181–184 (2004).
Han, J.W. et al. Genome-wide association study in a Chinese Han population identifies nine new susceptibility loci for systemic lupus erythematosus. Nat. Genet. 41, 1234–1237 (2009).
Zhang, X.J. et al. Psoriasis genome-wide association study identifies susceptibility variants within LCE gene cluster at 1q21. Nat. Genet. 41, 205–210 (2009).
Zhang, F.R. et al. Genomewide association study of leprosy. N. Engl. J. Med. 361, 2609–2618 (2009).
Campomenosi, P. et al. Characterization of RNASET2, the first human member of the Rh/T2/S family of glycoproteins. Arch. Biochem. Biophys. 449, 17–26 (2006).
Acquati, F. et al. Cloning and characterization of a senescence inducing and class II tumor suppressor gene in ovarian carcinoma at chromosome region 6q27. Oncogene 20, 980–988 (2001).
Monti, L. et al. RNASET2 as a tumor antagonizing gene in a melanoma cancer model. Oncol. Res. 17, 69–74 (2008).
Thompson, D.M. & Parker, R. The RNase Rny1p cleaves tRNAs and promotes cell death during oxidative stress in Saccharomyces cerevisiae. J. Cell Biol. 185, 43–50 (2009).
Schallreuter, K.U. et al. In vivo and in vitro evidence for hydrogen peroxide (H2O2) accumulation in the epidermis of patients with vitiligo and its successful removal by a UVB-activated pseudocatalase. J. Investig. Dermatol. Symp. Proc. 4, 91–96 (1999).
Mikolajka, A. et al. Structure of the N-terminal domain of the FOP (FGFR1OP) protein and implications for its dimerization and centrosomal localization. J. Mol. Biol. 359, 863–875 (2006).
Popovici, C. et al. The t(6;8)(q27;p11) translocation in a stem cell myeloproliferative disorder fuses a novel gene, FOP, to fibroblast growth factor receptor 1. Blood 93, 1381–1389 (1999).
Acquaviva, C. et al. The centrosomal FOP protein is required for cell cycle progression and survival. Cell Cycle 8, 1217–1227 (2009).
Schutyser, E., Struyf, S. & Van Damme, J. The CC chemokine CCL20 and its receptor CCR6. Cytokine Growth Factor Rev. 14, 409–426 (2003).
Le Borgne, M. et al. Dendritic cells rapidly recruited into epithelial tissues via CCR6/CCL20 are responsible for CD8+ T cell crosspriming in vivo. Immunity 24, 191–201 (2006).
Birlea, S.A., Gowan, K., Fain, P.R. & Spritz, R.A. Genome-wide association study of generalized vitiligo in an isolated European founder population identifies SMOC2, in close proximity to IDDM8. J. Invest. Dermatol. 130, 798–803 (2010).
Barrett, J.C. et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn's disease. Nat. Genet. 40, 955–962 (2008).
Levy, C. et al. Identifying a common molecular mechanism for inhibition of MITF and STAT3 by PIAS3. Blood 107, 2839–2845 (2006).
Garraway, L.A. et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 436, 117–122 (2005).
Imielinski, M. et al. Common variants at five new loci associated with early-onset inflammatory bowel disease. Nat. Genet. 41, 1335–1340 (2009).
Zhang, Z. et al. The analysis of genetics and associated autoimmune diseases in Chinese vitiligo patients. Arch. Dermatol. Res. 301, 167–173 (2009).
Taïeb, A. & Picardo, M. The definition and assessment of vitiligo: a consensus report of the Vitiligo European Task Force. Pigment Cell Res. 20, 27–35 (2007).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J. Natl. Cancer Inst. 22, 719–748 (1959).
Barrett, J.C., Fry, B., Maller, J. & Daly, M.J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
de Bakker, P.I. et al. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nat. Genet. 38, 1166–1172 (2006).
Browning, S.R. & Browning, B.L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
Acknowledgements
We thank all study participants. We also thank J.-J. Liu, J.-M. Chen and K. Seng Sim at the Genome Institute of Singapore for performing the imputation, association and LD analyses of HLA alleles. This study was supported in part by the Key Project of the Chinese National Natural Science Foundation (No. 30530670), the Cooperation Project of the Chinese Key National Natural Science Foundation for Overseas Youth (No. 30628021) and the Anhui Provincial Special Scientific Program (2007-7).
Author information
Authors and Affiliations
Contributions
X.-J.Z. conceived this study and obtained financial support; X.-J.Z., Y.Z. and S.Y. designed the study; C.Q., Y.-Q.R., L.-H.X. and L.-D.S. participated in the design and were responsible for sample selection, genotyping and project management; S.-M.Z., D.-Y.H., Yang Li, X.-F.T., S.-X.X., Y.-M.L., Q.X., J.-P.G., W.-L.H., C.N., T.-M.P., Yun Li, Y.-S.L. and Z.-Y.Y. conducted sample selection and managed data; A.-E.X., X.-H.G., H.-D.C., X.-M.P., R.-N.W., C.-Z.L., J.-B.L., T.-W.G., J.-Z.Z., X.-L.W., J.W., R.-Y.Y., L.L., J.-B.Y., Y.-W.L., X.-D.W., W.-S.H., Y.Y., Y.Z., W.-S.W., P.-L.D., K.L., X.-J.K., Y.-Q.L., L.S., Z.-F.L. and S.-Q.X. undertook recruitment, collected phenotype data, undertook related data handling and calculation and obtained biological samples; H.-Q.J., M.S., C.-Y.Z., Y.W., G.C., P.L., J.Z., H.C., M.L., X.Z., H.-Y.T., S.-M.H., S.Y. and C.-F.H. performed genotyping analyses; X.-B.Z., S.-Q.Z., X.-D.Z., X.-Y.Y. and F.-Y.Z. undertook data processing, statistical analysis and bioinformatics investigations; all the authors contributed to the final paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–4 (PDF 495 kb)
Rights and permissions
About this article
Cite this article
Quan, C., Ren, YQ., Xiang, LH. et al. Genome-wide association study for vitiligo identifies susceptibility loci at 6q27 and the MHC. Nat Genet 42, 614–618 (2010). https://doi.org/10.1038/ng.603
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.603
This article is cited by
-
Hashimoto’s thyroiditis and coexisting disorders in correlation with HLA status—an overview
Wiener Medizinische Wochenschrift (2023)
-
A comprehensive meta-analysis and prioritization study to identify vitiligo associated coding and non-coding SNV candidates using web-based bioinformatics tools
Scientific Reports (2022)
-
Genome-wide association study reveals ethnicity-specific SNPs associated with ankylosing spondylitis in the Taiwanese population
Journal of Translational Medicine (2022)
-
FBXO6-mediated RNASET2 ubiquitination and degradation governs the development of ovarian cancer
Cell Death & Disease (2021)
-
An in-depth analysis reveals two new genetic variants on 22q11.2 associated with vitiligo in the Chinese Han population
Molecular Biology Reports (2021)