Abstract
Purpose
Despite great advances that have been made in the understanding of the molecular complexity of acute myeloid leukemia (AML), very little has been translated into new therapies. Here, we set out to investigate the impact of cytoskeleton regulatory genes on clinical outcomes and their potential as therapeutic targets in AML.
Methods
Gene expression and clinical data were retrieved from The Cancer Genome Atlas (TCGA) AML study and used for survival and functional genomics analyses. For pharmacological tests, AML cells were exposed to ezrin (EZR) inhibitors and submitted to several cellular and molecular assays.
Results
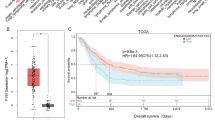
High EZR expression was identified as an independent marker of worse outcomes in AML patients from the TCGA cohort (p < 0.05). Functional genomics analyses suggested that EZR contributes to responses to stimuli and signal transduction pathways in leukemia cells. EZR pharmacological inhibition with NSC305787 and NSC668394 reduced viability, proliferation, autonomous clonal growth, and cell cycle progression in AML cells (p < 0.05). NSC305787 had a greater potency and efficiency than NSC668394 in leukemia models. At the molecular level, EZR inhibitors reduced EZR, S6 ribosomal protein and 4EBP1 phosphorylation, and induced PARP1 cleavage in AML cells. NSC305787, but not NSC668394, favored a gene network involving cell cycle arrest and apoptosis in Kasumi 1 AML cells.
Conclusions
From our data we conclude that EZR expression may serve as a prognostic factor in AML. Our preclinical findings indicate that ezrin inhibitors may be employed as a putative novel class of AML targeting drugs.





Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Code availability
Not applicable.
References
H. Dohner, D.J. Weisdorf, C.D. Bloomfield, Acute myeloid leukemia. N. Engl. J. Med. 373, 1136–1152 (2015)
X. Zhao, Y. Li, H. Wu, A novel scoring system for acute myeloid leukemia risk assessment based on the expression levels of six genes. Int. J. Mol. Med. 42, 1495–1507 (2018)
T.J. Ley, C. Miller, L. Ding, B.J. Raphael, A.J. Mungall, A. Robertson, K. Hoadley, T.J. Triche Jr., P.W. Laird, J.D. Baty, L.L. Fulton, R. Fulton, S.E. Heath, J. Kalicki-Veizer, C. Kandoth, J.M. Klco, D.C. Koboldt, K.L. Kanchi, S. Kulkarni, T.L. Lamprecht, D.E. Larson, L. Lin, C. Lu, M.D. McLellan, J.F. McMichael, J. Payton, H. Schmidt, D.H. Spencer, M.H. Tomasson, J.W. Wallis, L.D. Wartman, M.A. Watson, J. Welch, M.C. Wendl, A. Ally, M. Balasundaram, I. Birol, Y. Butterfield, R. Chiu, A. Chu, E. Chuah, H.J. Chun, R. Corbett, N. Dhalla, R. Guin, A. He, C. Hirst, M. Hirst, R.A. Holt, S. Jones, A. Karsan, D. Lee, H.I. Li, M.A. Marra, M. Mayo, R.A. Moore, K. Mungall, J. Parker, E. Pleasance, P. Plettner, J. Schein, D. Stoll, L. Swanson, A. Tam, N. Thiessen, R. Varhol, N. Wye, Y. Zhao, S. Gabriel, G. Getz, C. Sougnez, L. Zou, M.D. Leiserson, F. Vandin, H.T. Wu, F. Applebaum, S.B. Baylin, R. Akbani, B.M. Broom, K. Chen, T.C. Motter, K. Nguyen, J.N. Weinstein, N. Zhang, M.L. Ferguson, C. Adams, A. Black, J. Bowen, J. Gastier-Foster, T. Grossman, T. Lichtenberg, L. Wise, T. Davidsen, J.A. Demchok, K.R. Shaw, M. Sheth, H.J. Sofia, L. Yang, J.R. Downing, G. Eley, Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013)
F. Mollinedo, C. Gajate, Microtubules, microtubule-interfering agents and apoptosis. Apoptosis 8, 413–450 (2003)
E. Nogales, Structural insight into microtubule function. Annu. Rev. Biophys. Biomol. Struct. 30, 397–420 (2001)
M. Hesse, T.M. Magin, K. Weber, Genes for intermediate filament proteins and the draft sequence of the human genome: novel keratin genes and a surprisingly high number of pseudogenes related to keratin genes 8 and 18. J. Cell Sci. 114, 2569–2575 (2001)
I. Szeverenyi, A.J. Cassidy, C.W. Chung, B.T. Lee, J.E. Common, S.C. Ogg, H. Chen, S.Y. Sim, W.L. Goh, K.W. Ng, J.A. Simpson, L.L. Chee, G.H. Eng, B. Li, D.P. Lunny, D. Chuon, A. Venkatesh, K.H. Khoo, W.H. McLean, Y.P. Lim, E.B. Lane, The human intermediate filament database: comprehensive information on a gene family involved in many human diseases. Hum. Mutat. 29, 351–360 (2008)
A. Satelli, S. Li, Vimentin in cancer and its potential as a molecular target for cancer therapy. Cell. Mol. Life Sci. 68, 3033–3046 (2011)
K. Rottner, J. Faix, S. Bogdan, S. Linder, E. Kerkhoff, Actin assembly mechanisms at a glance. J. Cell Sci. 130, 3427–3435 (2017)
T. Fujii, A.H. Iwane, T. Yanagida, K. Namba, Direct visualization of secondary structures of F-actin by electron cryomicroscopy. Nature 467, 724–728 (2010)
T.Y. Liaw, M.H. Chang, M. Kavallaris, The cytoskeleton as a therapeutic target in childhood acute leukemia: obstacles and opportunities. Curr. Drug Targets 8, 739–749 (2007)
C. Thiele, G. Hirschfeld, cutpointr: Improved estimation and validation of optimal cutpoints in R. arXiv:2002.09209 (2020)
P.J. Heagerty, Y. Zheng, Survival model predictive accuracy and ROC curves. Biometrics 61, 92–105 (2005)
E. Cerami, J. Gao, U. Dogrusoz, B.E. Gross, S.O. Sumer, B.A. Aksoy, A. Jacobsen, C.J. Byrne, M.L. Heuer, E. Larsson, Y. Antipin, B. Reva, A.P. Goldberg, C. Sander, N. Schultz, The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012)
J. Gao, B.A. Aksoy, U. Dogrusoz, G. Dresdner, B. Gross, S.O. Sumer, Y. Sun, A. Jacobsen, R. Sinha, E. Larsson, E. Cerami, C. Sander, N. Schultz, Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013)
A.R. Lucena-Araujo, J.L. Coelho-Silva, D.A. Pereira-Martins, D.R. Silveira, L.C. Koury, R.A.M. Melo, R. Bittencourt, K. Pagnano, R. Pasquini, E.C. Nunes, E.M. Fagundes, A.B. Gloria, F. Kerbauy, M. de Lourdes Chauffaille, I. Bendit, V. Rocha, A. Keating, M.S. Tallman, R.C. Ribeiro, R. Dillon, A. Ganser, B. Lowenberg, P.J.M. Valk, F. Lo-Coco, M.A. Sanz, N. Berliner, E.M. Rego, Combining gene mutation with gene expression analysis improves outcome prediction in acute promyelocytic leukemia. Blood 134, 951–959 (2019)
W. Sauerbrei, M. Schumacher, A bootstrap resampling procedure for model building: application to the Cox regression model. Stat. Med. 11, 2093–2109 (1992)
A. Subramanian, P. Tamayo, V.K. Mootha, S. Mukherjee, B.L. Ebert, M.A. Gillette, A. Paulovich, S.L. Pomeroy, T.R. Golub, E.S. Lander, J.P. Mesirov, Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 102, 15545–15550 (2005)
T.L. Carrour, S. Assou, S. Tondeur, L. Lhermitte, N. Lamb, T. Reme, V. Pantesco, S. Hamamah, B. Klein, J. De Vos, Amazonia!: An online resource to google and visualize public human whole genome expression data. Open Bioinform. J. 4, 5–10 (2010)
H. Dohner, E. Estey, D. Grimwade, S. Amadori, F.R. Appelbaum, T. Buchner, H. Dombret, B.L. Ebert, P. Fenaux, R.A. Larson, R.L. Levine, F. Lo-Coco, T. Naoe, D. Niederwieser, G.J. Ossenkoppele, M. Sanz, J. Sierra, M.S. Tallman, H.F. Tien, A.H. Wei, B. Lowenberg, C.D. Bloomfield, Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood 129, 424–447 (2017)
K.J. Livak, T.D. Schmittgen, Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 (2001)
A.I. Saeed, V. Sharov, J. White, J. Li, W. Liang, N. Bhagabati, J. Braisted, M. Klapa, T. Currier, M. Thiagarajan, A. Sturn, M. Snuffin, A. Rezantsev, D. Popov, A. Ryltsov, E. Kostukovich, I. Borisovsky, Z. Liu, A. Vinsavich, V. Trush, J. Quackenbush, TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34, 374–378 (2003)
G. Bulut, S.H. Hong, K. Chen, E.M. Beauchamp, S. Rahim, G.W. Kosturko, E. Glasgow, S. Dakshanamurthy, H.S. Lee, I. Daar, J.A. Toretsky, C. Khanna, A. Uren, Small molecule inhibitors of ezrin inhibit the invasive phenotype of osteosarcoma cells. Oncogene 31, 269–281 (2012)
A. Ghaffari, V. Hoskin, G. Turashvili, S. Varma, J. Mewburn, G. Mullins, P.A. Greer, F. Kiefer, A.G. Day, Y. Madarnas, S. SenGupta, B.E. Elliott, Intravital imaging reveals systemic ezrin inhibition impedes cancer cell migration and lymph node metastasis in breast cancer. Breast Cancer Res. 21, 12 (2019)
H. Celik, S.H. Hong, D.D. Colon-Lopez, J. Han, Y.S. Kont, T.Z. Minas, M. Swift, M. Paige, E. Glasgow, J.A. Toretsky, J. Bosch, A. Uren, Identification of novel ezrin inhibitors targeting metastatic osteosarcoma by screening open access malaria box. Mol. Cancer Ther. 14, 2497–2507 (2015)
A. Corcoran, T.G. Cotter, FLT3-driven redox-modulation of Ezrin regulates leukaemic cell migration. Free Radic. Res. 47, 20–34 (2013)
R. Monni, L. Haddaoui, A. Naba, I. Gallais, M. Arpin, P. Mayeux, F. Moreau-Gachelin, Ezrin is a target for oncogenic Kit mutants in murine erythroleukemia. Blood 111, 3163–3172 (2008)
G. Habif, M.F. Grasset, S. Kieffer-Jaquinod, L. Kuhn, G. Mouchiroud, S. Gobert-Gosse, Phosphoproteome analyses reveal specific implications of Hcls1, p21-activated kinase 1 and Ezrin in proliferation of a myeloid progenitor cell line downstream of wild-type and ITD mutant Fms-like tyrosine kinase 3 receptors. J. Proteomics 78, 231–244 (2013)
K. Krishnan, B. Bruce, S. Hewitt, D. Thomas, C. Khanna, L.J. Helman, Ezrin mediates growth and survival in Ewing’s sarcoma through the AKT/mTOR, but not the MAPK, signaling pathway. Clin. Exp. Metastasis 23, 227–236 (2006)
E.H. Estey, Treatment of acute myeloid leukemia. Haematologica 94, 10–16 (2009)
K. Dohner, P. Paschka, Intermediate-risk acute myeloid leukemia therapy: current and future. Hematology Am. Soc. Hematol. Educ. Program 2014, 34–43 (2014)
Y. Ofran, J.M. Rowe, Genetic profiling in acute myeloid leukaemia–where are we and what is its role in patient management. Br. J. Haematol. 160, 303–320 (2013)
A. Bretscher, D. Reczek, M. Berryman, Ezrin: a protein requiring conformational activation to link microfilaments to the plasma membrane in the assembly of cell surface structures. J. Cell Sci. 110(Pt 24), 3011–3018 (1997)
K. Kawaguchi, S. Yoshida, R. Hatano, S. Asano, Pathophysiological roles of Ezrin/Radixin/Moesin proteins. Biol. Pharm. Bull. 40, 381–390 (2017)
A.L. Neisch, R.G. Fehon, Ezrin, Radixin and Moesin: key regulators of membrane-cortex interactions and signaling. Curr. Opin. Cell Biol. 23, 377–382 (2011)
N. Li, J. Kong, Z. Lin, Y. Yang, T. Jin, M. Xu, J. Sun, L. Chen, Ezrin promotes breast cancer progression by modulating AKT signals. Br. J. Cancer 120, 703–713 (2019)
Y. Song, X. Ma, M. Zhang, M. Wang, G. Wang, Y. Ye, W. Xia, Ezrin mediates invasion and metastasis in tumorigenesis: A review. Front. Cell. Dev. Biol. 8, 588801 (2020)
J. Li, K. Wei, H. Yu, D. Jin, G. Wang, B. Yu, Prognostic value of Ezrin in various cancers: A systematic review and updated meta-analysis. Sci. Rep. 5, 17903 (2015)
B. Zhou, J. Leng, M. Hu, L. Zhang, Z. Wang, D. Liu, X. Tong, B. Yu, Y. Hu, C. Deng, Y. Liu, Q. Zhang, Ezrin is a key molecule in the metastasis of MOLT4 cells induced by CCL25/CCR9. Leuk. Res. 34, 769–776 (2010)
H. Quentmeier, J. Reinhardt, M. Zaborski, H.G. Drexler, FLT3 mutations in acute myeloid leukemia cell lines. Leukemia 17, 120–124 (2003)
A. Beghini, I. Magnani, C.B. Ripamonti, L. Larizza, Amplification of a novel c-Kit activating mutation Asn(822)-Lys in the Kasumi-1 cell line: a t(8;21)-Kit mutant model for acute myeloid leukemia. Hematol. J. 3, 157–163 (2002)
B. Lowenberg, W.L. van Putten, I.P. Touw, R. Delwel, V. Santini, Autonomous proliferation of leukemic cells in vitro as a determinant of prognosis in adult acute myeloid leukemia. N. Engl. J. Med. 328, 614–619 (1993)
D. Pore, J. Bodo, A. Danda, D. Yan, J.G. Phillips, D. Lindner, B.T. Hill, M.R. Smith, E.D. Hsi, N. Gupta, Identification of Ezrin-Radixin-Moesin proteins as novel regulators of pathogenic B-cell receptor signaling and tumor growth in diffuse large B-cell lymphoma. Leukemia 29, 1857–1867 (2015)
H. Celik, G. Bulut, J. Han, G.T. Graham, T.Z. Minas, E.J. Conn, S.H. Hong, G.T. Pauly, M. Hayran, X. Li, M. Ozdemirli, A. Ayhan, M.A. Rudek, J.A. Toretsky, A. Uren, Ezrin inhibition up-regulates stress response gene expression. J. Biol. Chem. 291, 13257–13270 (2016)
Y. Tang, X. Sun, S. Yu, X. Bie, J. Wang, L. Ren, Inhibition of Ezrin suppresses cell migration and invasion in human nasopharyngeal carcinoma. Oncol. Lett. 18, 553–560 (2019)
X. Wan, A. Mendoza, C. Khanna, L.J. Helman, Rapamycin inhibits ezrin-mediated metastatic behavior in a murine model of osteosarcoma. Cancer Res. 65, 2406–2411 (2005)
C.H. Brandts, B. Sargin, M. Rode, C. Biermann, B. Lindtner, J. Schwable, H. Buerger, C. Muller-Tidow, C. Choudhary, M. McMahon, W.E. Berdel, H. Serve, Constitutive activation of Akt by Flt3 internal tandem duplications is necessary for increased survival, proliferation, and myeloid transformation. Cancer Res. 65, 9643–9650 (2005)
L. Larizza, I. Magnani, A. Beghini, The Kasumi-1 cell line: a t(8;21)-kit mutant model for acute myeloid leukemia. Leuk. Lymphoma 46, 247–255 (2005)
Y. Tabe, A. Tafuri, K. Sekihara, H. Yang, M. Konopleva, Inhibition of mTOR kinase as a therapeutic target for acute myeloid leukemia. Expert. Opin. Ther. Targets 21, 705–714 (2017)
S. Darici, H. Alkhaldi, G. Horne, H.G. Jorgensen, S. Marmiroli, X. Huang, Targeting PI3K/Akt/mTOR in AML: Rationale and clinical evidence. J. Clin. Med. 9, 2934 (2020)
I. Nepstad, K.J. Hatfield, I.S. Gronningsaeter, H. Reikvam, The PI3K-Akt-mTOR signaling pathway in human acute myeloid leukemia (AML) cells. Int. J. Mol. Sci. 21, 2907 (2020)
M. Ingham, G.K. Schwartz, Cell-cycle therapeutics come of age. J. Clin. Oncol. 35, 2949–2959 (2017)
M.P. Luna-Vargas, J.E. Chipuk, The deadly landscape of pro-apoptotic BCL-2 proteins in the outer mitochondrial membrane. FEBS J. 283, 2676–2689 (2016)
W.A. Siddiqui, A. Ahad, H. Ahsan, The mystery of BCL2 family: Bcl-2 proteins and apoptosis: an update. Arch. Toxicol. 89, 289–317 (2015)
H.E. Ramsey, M.A. Fischer, T. Lee, A.E. Gorska, M.P. Arrate, L. Fuller, K.L. Boyd, S.A. Strickland, J. Sensintaffar, L.J. Hogdal, G.D. Ayers, E.T. Olejniczak, S.W. Fesik, M.R. Savona, A novel MCL1 inhibitor combined with Venetoclax rescues Venetoclax-resistant acute myelogenous leukemia. Cancer Discov. 8, 1566–1581 (2018)
Acknowledgements
The authors thank Dr. Gabriela Sarti Kinker for providing assistance with the GSEA analysis. The authors also would like to acknowledge all the research participants contributing to The Cancer Genome Atlas (TCGA) resource for providing high quality data for analysis.
Funding
This study was supported by grants #2017/24993-0, #2019/23864-7 and #2015/17177-6 from the São Paulo Research Foundation (FAPESP). This study was also financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001.
Author information
Authors and Affiliations
Contributions
J.C.L.S. conception and design, execution of experiments, data analysis and interpretation, and manuscript writing, J.L.C.-S., H.P.V. and K.L. conception and design, data analysis and interpretation, and manuscript editing. M.L., L.V.C.-L. and F.T. data analysis and interpretation, and manuscript writing. J.A.M.-N. conception and design, data analysis and interpretation, manuscript writing and gave final approval of the manuscript. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Conflict of interest disclosure
The authors declare no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Lipreri da Silva, J.C., Coelho-Silva, J.L., Lima, K. et al. Comprehensive analysis of cytoskeleton regulatory genes identifies ezrin as a prognostic marker and molecular target in acute myeloid leukemia. Cell Oncol. 44, 1105–1117 (2021). https://doi.org/10.1007/s13402-021-00621-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-021-00621-0