Abstract
Background
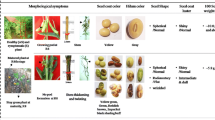
The plants of B. rapa (syn. B. campestris) are the most important food crop of Pakistan for the production of cooking oil. Brassica plants infected by phytoplasma exhibit floral abnormalities including phyllody, virescence, hypertrophied sepal and aborted reproductive organs and affected flower developmental genes which reduces the yield manifold.
Methods and results
The expression level of flower developmental genes in healthy and phytoplasma infected brassica were compared by using semi-quantitative reverse transcription polymerase chain reaction and DNA hybridization. In infected brassica, LEAFY (LFY) gene, controlling the development and maintenance of floral organ, and directly involved in controlling the homeotic gene expression was affected, while APETALA2, regulate the production of sepals and petals, were not altered. Whereas the genes WUSCHEL, APETALA3 and AGAMOUS, were significantly down-regulated, that were responsible for the identity of shoot and central meristem, petals and stamens production, and stamens and carpels development, respectively. The GLUB gene, controlling the production of β-1,3-glucanases enzyme, was highly up-regulated. According to DNA hybridization results, AGAMOUS and APETALA3 were restricted to floral organs territories in healthy and phytoplasma infected brassica, indicating that their expression was tissue-specific. These outcomes indicated that flower abnormalities of phytoplasma infected B. rapa are linked with DNA methylation in the expression of homeotic genes regulating flower development.
Conclusions
Azacitidine act as a DNA demethylating reagent. By applying the foliar spray of azacitidine during the flower development, cells of Phytoplasma infected plants exhibits demethylation of DNA when treated with azacitidine chemical that incorporated as analogue of cytosine during the cell division stage. B. rapa showed the up-regulation of gene expression level significantly that restore the normal production of flowers, ultimately increase the oil production throughout the world.





Similar content being viewed by others
Change history
15 November 2022
A Correction to this paper has been published: https://doi.org/10.1007/s11033-022-07912-1
References
Scarth R, Tang T (2006) Modification of brassica oil using conventional and transgenic approaches. Health Sci 41:67–71
Liefting LW, Shaw M, Kirkpatrick BC (2004) Sequence analysis of two plasmids from the phytoplasma beet leafhopper transmitted virescence agent. Microbiology 150:1809–1817
Hogenhout, (2011) Diverse targets of phytoplasma effectors: from plant development to defense against insects. Annu Rev Phytopathol 49:175–195
Calari A, Paltrinieri S, Contaldo N, Sakalieva D, Mori N, Duduk B, Bertaccini A (2011) Molecular evidence of phytoplasmas in winter oilseed rape, tomato and corn seedlings. Bull Insectology 64:157–158
Davis RE, Lee IM (2000) Phytoplasma. In: Lederberg J (ed) Encyclopedia of microbiology, 2nd edn. Academic Press, Cambridge, pp 640–646
Ahmad JN, Sharif MZ, Ahmad SJN, Tahir M, Bertaccini A (2019) Molecular identification and characterization of phytoplasmas in insect vectors of chickpea phyllody disease in Punjab Pakistan. Phytopathogenic Mollicutes 9(1):105–106
Sharif MZ, Ahmad SJN, Tahir M, Ziaf K, Zhang SH, Ahmad JN (2019) Molecular identification and characterization of phytoplasmas associated with carrot, cabbage and onion crops and their insect vectors in Punjab Pakistan. Pak J Agric Sci 56(2):1–8
Jung HY, Sawayanagi T, Wongkaew P, Miyata S, Ugaki M, Hibi T, Namba S (2003) ‘Candidatus phytoplasma oryzae’, a novel phytoplasma taxon associated with rice yellow dwarf disease. Int J Syst Evol Microbiol 53:1925–1929
Arocha Y, Lopez M, Pinol B, Fernandez M, Picornell B, Almeida R, Palenzuela WMR, Jones P (2005) ‘Candidatus phytoplasma graminis’ and ‘Candidatus phytoplasma caricae’, two novel phytoplasmas associated with diseases of sugarcane, weeds and papaya in Cuba. Int J Syst Evol Microbiol 55:2451–2463
Seemuller E, Schneider B (2004) ‘Candidatus phytoplasma mali’, ‘Candidatus phytoplasma pyri’ and ‘Candidatus phytoplasma prunorum’, the causal agents of apple proliferation, pear decline and European stone fruit yellows, respectively. Int J Syst Evol Microbiol 4:1217–1226
Al-Saady NA, Akhtar JK, Alberto C, Ali MS, Assunta B (2008) ‘Candidatus phytoplasma omanense’, associated with witches’-broom of Cassia italica (Mill.) Spreng. in Oman. Int J Syst Evol Microbiol 58:461–466
Valiunas D, Staniulis J, Davis RE (2006) ‘Candidatus Phytoplasma fragariae’, a novel phytoplasma taxon discovered in yellows diseased strawberry, Fragaria x ananassa. Annu Rev Phytopathol 56:277–281
Bertaccini A (2007) Phytoplasmas: diversity, taxonomy and epidemiology. Front Biol 12:673–689
Schneider B, Torres E, Martin MP, Schroder M, Behnke HD, Seemuller E (2005) ‘Candidatus phytoplasma pini’, a novel taxon from Pinus silvestris and Pinus halpensis. Int J Syst Evol Microbiol 55:303–307
Curkovic-Perica M, Lepedus H, Seruga-Music M (2007) Effect of indole-3-butyric acid on phytoplasmas in infected Cathan anthusroseus shoots grown in vitro. FEMS Microbiol Lett 268:171–177
Olivier CY, Galka B (2008) Consequences of phytoplasma infection on canola crop production in the Canada prairies. In: Endure International Conference, Diversifying crop protection, La grande-Motte, France, Book of Abstracts, pp 1–4
Verdin E, Pascal S, Jean-Luc D, Elia C, Fouad J, Souheir EZ, Brigitte G, Joseph MB, Monique G (2003) ‘Candidatus phytoplasma phoenicium’ sp. nov., a novel phytoplasma associated with an emerging lethal disease of almond trees in Lebanon and Iran. Annu Rev Phytopathol 53:833–838
Adwas A, Elsayed A, Azab A (2019) Oxidative stress and antioxidant mechanisms in human body. J Appl Biotechnol Bioeng 6(1):43–47
Firrao G, Gibb K, Streten C (2005) Short taxonomic guide to the genus ‘Candidatus Phytoplasma’. Plant Pathol 87:249–263
Himeno M, Neriya Y, Minato N, Miura C, Sugawara K, Ishii Y, Amaji Y, Kakizawa S, Oshima K, Namba S (2011) Unique morphological changes in plant pathogenic phytoplasma-infected petunia flowers are related to transcriptional regulation of floral homeotic genes in an organ-specific manner. Plant J 67:971–979
Kitamura Y, Hosokawa M, Uemachi U, Yazawa S (2009) Selection of ABC genes for candidate genes of morphological changes in hydrange a floral organs induced by phytoplasma infection. Sci Hortic 122:603–609
Pracros P, Renaudin J, Eveillard S, Mouras A, Hernould M (2006) Tomato flower abnormalities induced by stolbur phytoplasma infection are associated with changes of expression of floral development genes. Mol Plant Microbe Interact 19:62–68
Reeves PH, Coupland G (2001) Analysis of flowering time control in arabidopsis by comparison of double and triple mutants. Plant Physiol 126:1085–1091
Sugio A, Maclean AM, Kingdom HN, Grieve VM, Manimekalai R, Hogenhout SA (2011) Diverse targets of phytoplasma effectors: from plant development to defense against insects. Annu Rev Phytopathol 49:175–195
Smaczniak C, Immink RG, Angenent GC, Kaufmann K (2012) Developmental and evolutionary diversity of plant MADS-domain factors: insights from recent studies. Development 139(17):3081–3098
Ahmad JN, Garcion C, Teyssier T, Hernould M, Gallusci P, Pracros P, Renaudin J, Evellerd S (2013) Effects of stolbur phytoplasma infection on DNA methylation processes in tomato plants. Plant Pathol 62:205–216
Prunet N, Morel P, Negrutiu I, Trehin C (2009) Time to stop: flower meristem termination. Plant Physiol 150:1764–1772
Su YT, Chen J-C, Lin C-P (2011) Phytoplasma-induced floral abnormalities in catharanthus roseus are associated with phytoplasma accumulation and transcript repression of floral organ identity genes. Mol Plant Microbe Interact 24:1502–1512
Zheng X, Chen L, Li M, Lou Q, Xia H, Wang P, Li T, Liu H, Luo L (2013) Transgenerational variations in DNA methylation induced by drought stress in two rice varieties with distinguished difference to drought resistance. PLoS ONE 8:e80253
Mason G, Noris E, Lanteri S, Acquadro A, Accotto PG, Portis E (2008) Potentiality of methylation-sensitive amplification polymorphism (MSAP) in identifying genes involved in tomato response to tomato yellow leaf curl sardinia virus. Plant Mol Biol Rep 26:156–173
Cao X, Fan G, Deng M, Zhao Z, Dong Y (2014) Identification of genes related to paulownia witches’ broom by AFLP and MSAP. Int J Mol Sci 15:14669–14683
Jacobsen SE, Sakai H, Finnegan EJ, Cao X, Meyerowitz EM (2000) Ectopic hyper-methylation of flower-specific genes in Arabidopsis. Curr Biol 10:179–186
Yia H, Lihong L (2013) DNA methylation changes in response to sulfur dioxide stress in Arabidopsis plants. Procedia Environ Sci 18:37–42
Manning K, Tor M, Poole M et al (2006) A naturally occurring epigenetic mutation in a gene encoding an SBP-box transcription factor inhibits tomato fruit ripening. Nat Genet 38:948–952
Takeno K (2010) Epigenetic regulation of photoperiodic flowering. Plant Signal Behav 5:788–791
Ahrens U, Seemuller E (1992) Detection of DNA of plant pathogenic mycoplasma-like organisms by a polymerase chain reaction that amplifies a sequence of the 16S rRNA gene. Phytopathology 82:828
Foerster AM, Scheid OM (2010) Analysis of DNA methylation in plants by bisulfite sequencing. In: Kovalchuk I, Zemp FJ (eds) Plant Epigenetics. Humana Press, Totowa NJ USA, pp 1–11
Teyssier E, Bernacchia G, Maury S et al (2008) Tissue dependent variations of DNA methylation and endoreduplication levels during tomato fruit development and ripening. Planta 228:391–399
Mazzucato A, Olimpieri I, Siligato F, Picarella ME, Soressi GP (2008) Characterization of genes controlling stamen identity and development in a parthenocarpic tomato mutant indicates a role for the DEFICIENS ortholog in the control of fruit set. Physiol Plant 132:526–537
Lohmann JU, Weigel D (2002) Building beauty: the genetic control of floral patterning. Dev Cell 2:135–142
Ahmad JN, Renaudin J, Eveillard S (2014) Expression of defence genes in stolbur phytoplasma infected tomatoes, and effect of defense stimulators on disease development. Eur J Plant Pathol 139:39–51
MacLean AM, Sugio A, Makarova OV et al (2011) Phytoplasma effector SAP54 induces indeterminate leaf-like flower development in Arabidopsis plants. Plant Physiol 157:831–841
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16:6–21
Zilberman D, Gehring M, Tran RK, Ballinger T, Henikoff S (2007) Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat Genet 39(61):9–629
Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1992) LEAFY controls floral meristem identity in Arabidopsis. Cell 69:843–859
Lenhard M, Bohnert A, Jurgens G, Laux T (2001) Termination of stem cell maintenance in Arabidopsis floral meristems by interactions between WUSCHEL and AGAMOUS. Cell 105:805–814
Yanofsky MF, Ma H, Bowman JL, Drews GN, Feldmann KA, Meyerowitz EM (1990) The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors. Nature 34:35–39
Kaufmann K, Melzer R, Theissen G (2005) MIKC-type MADS domain proteins: structural modularity, protein interactions and network evolution in land plants. Gene 347:183–198
Robles P, Pelaz S (2005) Flower and fruit development in Arabidopsis thaliana. Int J Dev Biol 49:633–643
Cettul E, Firrao G (2010) Effects of phytoplasma infection on Arabidopsis thaliana development. In: Brown DR, Bertaccini A (eds) 18th Congress of the international organization for mycoplasmology chiancianoterme. Italy
Sharma V (2013) Pathogenesis related defence functions of plant chitinases and β-1,3-glucanases. Vegetos 26:215–218
Cordero MJ, Raventos D, San Segundo B (1994) Differential expression and induction of chitinases and β-1,3-glucanases in response to fungal infection during germination of maize seeds. Mol Plant Microbe Interact 7:23–31
Khan MA, Jafri S, Rana IA, Ullah I (1992) Improvement of wheat protein by supplementation with potato flour. Pak J Agric Sci 13:101–106
Ward ER, Payne GB, Moyer MB, Williams SC, Dincher SS, Sharkey KC, Beck JJ, Taylor HT, Ahl-Goy P, Meins FJ, Ryals JA (1991) Differential regulation of b-1,3-glucanase messenger RNAs inresponse to pathogen infection. Plant Physiol 96:390–397
Villanea F, Bolnick D, Monroe C, Worl R, Cambra R, Leventhal A (2013) Brief communication: evolution of a specific O allele (O1v (G542A)) supports unique ancestry of Native Americans. Am J Phys Anthropol 151:649–657
Llamas B, Holland ML, Chen K, Cropley JE, Cooper A, Suter CM (2012) High-resolution analysis of cytosine methylation in ancient DNA. PLoS ONE 7:1–6
Jofuku KD, Den-Boer BGW, Van-Montagu M, Okamuro JK (1994) Control of Arabidopsis flower and seed development by the homeotic gene APETALA2. Plant Cell 6:1211–1225
Zhang M, Kimatu JN, Xu K, Liu B (2010) DNA cytosine methylation in plant development. J Genet Genom 37:1–12
Buoso S, Pagliari L, Musetti R, Martini M, Marroni F, Schmidt W, Santi S (2019) ‘Candidatus Phytoplasma solani’interferes with the distribution and uptake of iron in tomato. BMC Genomics 20(1):703
Acknowledgements
The FICP international research grant run by Principal Investigator Dr. Jam Nazeer Ahmad and Co- principal investigator Dr. Samina Jam Nazeer Ahmad for research funding. All the equipment and chemical I used for my research work were funded by this project. The authors extend their appreciation to the researchers supporting project no (RSP-2021/293) at King Saud University, Riyadh, Saudi Arabia.
Funding
The authors have not disclosed any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to Dr. Hamada AbdElgawad's affiliation update and incomplete Acknowledgement section.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ahmad, M.A., Ahmad, S.J.N., Shah, A.N. et al. Study of genetic modifications of flower development and methylation status in phytoplasma infected Brassica (Brassica rapa L.). Mol Biol Rep 49, 11359–11369 (2022). https://doi.org/10.1007/s11033-022-07743-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-022-07743-0