Abstract
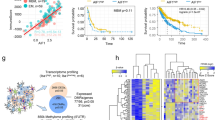
The immune cells of tumor microenvironment (TME) constitute a vital element of the tumor tissue. There is increasing evidence for their clinical significance in predicting prognosis and therapeutic outcomes. However, the TME immune cell infiltrating pattern of the bone marrow in acute myeloid leukemia (AML) patients remains unclear. Here, RNA-sequencing results of AML patients from TCGA database were used to quantify the abundance of 28 types of immune cells in the TME using the single-sample gene set enrichment analysis algorithm. We comprehensively evaluated the immune infiltration status in the TCGA-LAML cohort and defined two immunophenotypes: the immune hot and immune cold subtypes. Additionally, we constructed a TME score reflecting the immune infiltrating pattern of the patients using Cox regression algorithm. Subtypes with high TME score were characterized by over-activation of immune inflammation-related pathways, release of inflammatory factors, T-cell dysfunction, and poor prognosis. Subtypes with a low TME score were characterized by relatively low immune infiltration and immune exclusion. Our analysis indicated that patients in the low TME score group were more sensitive to chemotherapeutic drugs, and those in high TME score were more likely to respond to immunotherapy. Our study provides a new direction to evaluate anti-tumor therapy from immune infiltration of the TME, and the individualized scoring system in this study has important clinical significance in identifying patients who respond to immunotherapy.



Similar content being viewed by others
Data availability
The dataset TCGA-LAML analyzed during the current study is available in the Cancer Genome Atlas (TCGA) database (https://portal.gdc.cancer.gov/projects/TCGA-LAML). And the dataset GSE106291 analyzed during the current study is available in the Expression Omnibus (GEO) database under accession ID GSE106291 (http://www.ncbi.nlm.nih.gov/geo/). Associated codes are available at https://github.com/lwx0727/code-available.git.
References
Döhner H, Weisdorf DJ, Bloomfield CD. Acute myeloid leukemia. N Engl J Med. 2015;373(12):1136–52.
De Kouchkovsky I, Abdul-Hay M. Acute myeloid leukemia: a comprehensive review and 2016 update. Blood Cancer J. 2016;6(7):e441.
Medeiros BC, Chan SM, Daver NG, Jonas BA, Pollyea DA. Optimizing survival outcomes with post-remission therapy in acute myeloid leukemia. Am J Hematol. 2019;94(7):803–11.
Daver N, Garcia-Manero G, Basu S, et al. Efficacy, safety, and biomarkers of response to Azacitidine and Nivolumab in relapsed/refractory acute myeloid leukemia: a nonrandomized, open-label. Phase II Study Cancer Discov. 2019;9(3):370–83.
Goswami M, Gui G, Dillon LW, et al. (2022) Pembrolizumab and decitabine for refractory or relapsed acute myeloid leukemia. J Immunother Cancer. 10(1).
Zeidan AM, Cavenagh J, Voso MT, et al. Efficacy and safety of azacitidine (AZA) in combination with the anti-PD-L1 Durvalumab (durva) for the front-line treatment of older patients (pts) with acute myeloid leukemia (AML) who are unfit for intensive chemotherapy (IC) and Pts with higher-risk myelodysplastic syndromes (HR-MDS): results from a large, international, randomized phase 2 study. Blood. 2019;134:829–829.
Casey SC, Amedei A, Aquilano K, et al. Cancer prevention and therapy through the modulation of the tumor microenvironment. Semin Cancer Biol. 2015;35:S199–223.
Binnewies M, Roberts EW, Kersten K, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med. 2018;24(5):541–50.
Pitt JM, Marabelle A, Eggermont A, Soria JC, Kroemer G, Zitvogel L. Targeting the tumor microenvironment: removing obstruction to anticancer immune responses and immunotherapy. Ann Oncol Off J Eur Soc Med Oncol. 2016;27(8):1482–92.
Fridman WH, Zitvogel L, Sautès-Fridman C, Kroemer G. The immune contexture in cancer prognosis and treatment. Nat Rev Clin Oncol. 2017;14(12):717–34.
Kalluri R. The biology and function of fibroblasts in cancer. Nat Rev Cancer. 2016;16(9):582–98.
Mantovani A, Marchesi F, Malesci A, Laghi L, Allavena P. Tumour-associated macrophages as treatment targets in oncology. Nat Rev Clin Oncol. 2017;14(7):399–416.
Turley SJ, Cremasco V, Astarita JL. Immunological hallmarks of stromal cells in the tumour microenvironment. Nat Rev Immunol. 2015;15(11):669–82.
Quail DF, Joyce JA. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013;19(11):1423–37.
Ali HR, Chlon L, Pharoah PD, Markowetz F, Caldas C. Patterns of immune infiltration in breast cancer and their clinical implications: a gene-expression-based retrospective study. PLoS Med. 2016;13(12):e1002194.
Brück O, Dufva O, Hohtari H, et al. Immune profiles in acute myeloid leukemia bone marrow associate with patient age, T-cell receptor clonality, and survival. Blood Adv. 2020;4(2):274–86.
Vago L, Gojo I. Immune escape and immunotherapy of acute myeloid leukemia. J Clin Investig. 2020;130(4):1552–64.
Weinstein JN, Collisson EA, Mills GB, et al. The cancer genome atlas pan-cancer analysis project. Nat Genet. 2013;45(10):1113–20.
Herold T, Jurinovic V, Batcha AMN, et al. A 29-gene and cytogenetic score for the prediction of resistance to induction treatment in acute myeloid leukemia. Haematologica. 2018;103(3):456–65.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013;14:7.
Charoentong P, Finotello F, Angelova M, et al. Pan-cancer immunogenomic analyses reveal genotype-immunophenotype relationships and predictors of response to checkpoint blockade. Cell Rep. 2017;18(1):248–62.
Wilkerson MD, Hayes DN. ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics (Oxford, England). 2010;26(12):1572–3.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
Speiser JL, Miller ME, Tooze J, Ip E. A comparison of random forest variable selection methods for classification prediction modeling. Expert Syst Appl. 2019;134:93–101.
Bolstad BM, Irizarry RA, Astrand M, Speed TP. A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics (Oxford, England). 2003;19(2):185–93.
Tibshirani R. The lasso method for variable selection in the cox model. Stat Med. 1997;16(4):385–95.
Laska E, Meisner M, Wanderling J. A maximally selected test of symmetry about zero. Stat Med. 2012;31(26):3178–91.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7.
Subramanian A, Tamayo P, Mootha VK, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–50.
Geeleher P, Cox N, Huang RS. pRRophetic: an R package for prediction of clinical chemotherapeutic response from tumor gene expression levels. PLoS ONE. 2014;9(9):e107468.
Jiang P, Gu S, Pan D, et al. Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med. 2018;24(10):1550–8.
Korn C, Méndez-Ferrer S. Myeloid malignancies and the microenvironment. Blood. 2017;129(7):811–22.
Krause DS, Fulzele K, Catic A, et al. Differential regulation of myeloid leukemias by the bone marrow microenvironment. Nat Med. 2013;19(11):1513–7.
Szczepanski MJ, Szajnik M, Czystowska M, et al. Increased frequency and suppression by regulatory T cells in patients with acute myelogenous leukemia. Clin Cancer Res Off J Am Assoc Cancer Res. 2009;15(10):3325–32.
Lim HX, Kim TS, Poh CL. Understanding the differentiation, expansion, recruitment and suppressive activities of myeloid-derived suppressor cells in cancers. Int J Mol Sci. 2020;21(10):3599.
Lin Y, Xu J, Lan H. Tumor-associated macrophages in tumor metastasis: biological roles and clinical therapeutic applications. J Hematol Oncol. 2019;12(1):76.
Beyar-Katz O, Gill S. Novel Approaches to Acute Myeloid Leukemia Immunotherapy. Clin Cancer Res Off J Am Assoc Cancer Res. 2018;24(22):5502–15.
Isidori A, Salvestrini V, Ciciarello M, et al. The role of the immunosuppressive microenvironment in acute myeloid leukemia development and treatment. Expert Rev Hematol. 2014;7(6):807–18.
Chen DS, Mellman I. Elements of cancer immunity and the cancer-immune set point. Nature. 2017;541(7637):321–30.
Wang A, Zhong H. Roles of the bone marrow niche in hematopoiesis, leukemogenesis, and chemotherapy resistance in acute myeloid leukemia. Hematology (Amsterdam, Netherlands). 2018;23(10):729–39.
Tabe Y, Konopleva M. Role of microenvironment in resistance to therapy in AML. Curr Hematol Malig Rep. 2015;10(2):96–103.
Dufva O, Pölönen P, Brück O, et al. Immunogenomic landscape of hematological malignancies. Cancer Cell. 2020;38(3):380-399.e313.
Mehtonen J, Pölönen P, Häyrynen S, et al. Data-driven characterization of molecular phenotypes across heterogeneous sample collections. Nucl Acids Res. 2019;47(13):e76.
Acknowledgements
We sincerely appreciated the data provided by GEO and TCGA database.
Funding
The present study was funded by the National Natural Science Foundation of China (Grant No.: 82070133).
Author information
Authors and Affiliations
Contributions
All authors were responsible for the conception and design of the manuscript. WXL and GPY contributed to draft the manuscript. GPY, WXL, XFC, YLL, JXL, YXC, ZXZ, YJL, XYZ, and CXY contributed to develop the methodology, analyze the data, and interpret the results. DX was responsible for preparing the figures, acquiring the funding, and revising critically to the manuscript. All authors have read and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lu, W., Yu, G., Li, Y. et al. Identifying prognostic biomarker related to immune infiltration in acute myeloid leukemia. Clin Exp Med 23, 4553–4562 (2023). https://doi.org/10.1007/s10238-023-01164-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-023-01164-4