Abstract
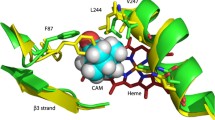
The effect of 1R-camphor on the conformational stability of the heme active site of cytochrome P450cam has been investigated. The absorption spectra of the heme moiety showed the presence of two hitherto unknown intermediates formed at low urea concentrations or during small temperature perturbations. The corresponding thermodynamic parameters were obtained by global fitting of the experimental data to a generalized sequential unfolding model at different wavelengths, which showed that the active conformation of the enzyme is stabilized by binding of the substrate at the active site. Circular-dichroism spectra of the enzyme in the visible- and far-UV region were studied to identify the critical range of denaturant concentration and the temperature at which the tertiary structure around the heme center was affected with almost no change in the secondary structure of the enzyme. This critical range of urea concentration was 0–2.8 M in the presence of camphor and 0–1.5 M in the absence of camphor. The tertiary structure of the enzyme was found to undergo conformational change in the temperature range 20–60 °C in the presence of the substrate and 20–47 °C in its absence. The spectral assignments of the intermediate species of the heme active site with the intact secondary structure of the enzyme were made by deconvolution of the Soret absorption spectra, and the results were analyzed to determine stabilization of the heme active-site geometry by 1R-camphor. Results showed that subtle conformational changes due to melting of the tertiary contacts in the active site lead to formation of intermediates which are coordinatively similar to the native enzyme. Analogous intermediate species might be responsible for leakage in the redox catalytic cycle of the enzyme.







Similar content being viewed by others
References
Mueller EJ, Loida PJ, Sligar SG (1995) In: Oritz de Montellano PR (ed) Cytochrome P450: structure, mechanism and biochemistry. Plenum Press, New York, pp 83–124
Wong L-L, Westlake ACG, Nickerson DP (1997) Struct Bonding 88:175–207
Oritz de Montellano PR (ed) (1995) Cytochrome P450: structure, mechanism and biochemistry. Plenum Press, New York, pp 83–123
Poulos TL, Finzel B, Howard AJ (1987) J Mol Biol 195:687–700
Omura T, Sato R (1964) J Biol Chem 239:2379–2385
Yu C- A, Gunsalus IC (1974) J Biol Chem 249:94–104
Poulos TL, Barry C, Finzel B, Gunsalus IC, Wagner GC, Kraut J (1985) J Biol Chem 260:16122–16130
Ogliaro F, de Visser SP, Cohen S, Sharma PK, Shaik S (2001) J Am Chem Soc 124:2806–2816
Schlichting I, Berendzen J, Chu K, Stock AM, Maves SA, Benson DE, Sweet RM, Ringe D, Petsko GA, Sligar SG (2000) Science 287:1615–1622
Ludemann SK, Carugo O, Wade RC (1997) J Mol Modell 3:369–374
Prasad S, Mazumdar S, Mitra S (2000) FEBS Lett 477:157–160
Peterson JA (1971) Arch Biochem Biophys 144:678–693
Peterson JA, Ishimura Y, Griffin BW (1972) Arch Biochem Biophys 149:197–208
Lange R, Pierre J, Debey P (1980) Eur J Biochem 107:441–445
Winn JW, Ludemann SK, Gauges R, Lounnas V, Wade RC (2002) Proc Natl Acad Sci USA 99:5361–5266
Deprez E, Gerber NC, Di Primo C, Douzou P, Sligar SG, Hui Bon Hoa G (1994) Biochemistry 33:14464–14468
Lange R, Hui Bon Hoa G, Debey P, Gunsalus IC (1977) Eur J Biochem 77:479–485
Sculze H, Hui Bon Hoa G, Helms V, Wade RC, Jung C (1996) Biochemistry 35:14127–14138
Gunsalus IC, Wagner GC (1978) Methods Enzymol 52:166–188
Jung C, Bendzko P, Ristau O, Gunsalus IC (1985) In: Vereczky L, Magyar K (eds) Cytochrome P450: biochemistry, biophysics and induction. Akademiai Kiado, Budapest, pp 19–22
Pfeil W, Nolting BO, Jung C (1993) Biochemistry 32:8856–8862
Nolting B, Jung C, Snatzke G (1992) Biochim Biophys Acta 1100:171–176
Mouro C, Jung C, Bondon A, Simonneaux G (1997) Biochemistry 36:8125–8134
Pfeil W, Nölting B, Jung C (1993) In: van den Tweel W, Harder A, Buitelaar R M (eds) Stability and stabilization of enzymes. Proc Int Symp, Studies Org Chem, Maastricht, The Netherlands, 22–25 November 1992. Elsevier, Amsterdam, pp 407–414
Jung C, Pfeil W, Köpke K, Schulze H, Ristau O (1994) In: Lechner MC (ed) Cytochrome P450, 8th Int Conf, Paris. pp 543–545
Hui Bon Hoa G, Marden MC (1982) Eur J Biochem 124:311–315
Di Primo C, Hui Bon Hoa G, Douzou P, Sligar SG (1992) Eur J Biochem 209:583–588
Marden MC, Hui Bon Hoa G (1982) Eur J Biochem 129:111–117
Yu X-C, Shen S, Strobel HW (1995) Biochemistry 34:5511–5517
Yoshioka S, Takahashi S, Hori H, Ishimori K, Morishima I (2001) Eur J Biochem 268:252–259
Auclair K, Moenne-Locoz P, Oritz de Montellano PR (2001) J Am Chem Soc 123:4877–4885
Unger BP, Gunsalus IC (1986) J Biol Chem 261:1158–1163
Press WH, Flanery BP, Teukolsky SA, Vetterling WT (1986) In: Numerical recipes. Cambridge University Press, Cambridge, p 523
Antonini E, Brunori M (1971) Hemoglobin and myoglobin in their reactions with ligands. North-Holland, Amsterdam
Martinis SA, Blanke SR, Hager LP, Sligar SG (1996) Biochemistry 35:14530–14536
Acknowledgements
This work was supported by the Tata Institute of Fundamental Research. Authors wish to thank Dr. L.L. Wong, University of Oxford, and Prof. S.G. Sligar, University of Illinois, Urbana-Champaign, for kindly providing E. coli strains of recombinant P450cam. Authors thank the referees for their constructive comments. RM thanks the TIFR Endowment Fund for Career Development for their support and Mr. Ram Reddy for his help in DLS experiments.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Let us assume a general unfolding model with n sequential (reversible) intermediates. The scheme can be written as follows:
where N denotes native protein, U denotes unfolded form, I is concentration of the respective intermediates, k values are corresponding rate constants, and K i (mol−1) is the equilibrium constant for the ith step. The coupled linear equation for the aforementioned scheme can be written as:
where \( I_{i} = {\left( {{k_{i} } \mathord{\left/ {\vphantom {{k_{i} } {k_{{ - i}} }}} \right. \kern-\nulldelimiterspace} {k_{{ - i}} }} \right)} \times I_{{i - 1}} = K_{i} I_{{i - 1}} \) and \( {\sum\nolimits_{i = 1}^{n + 1} {I_{i} = } }{\left( {{I_{1} } \mathord{\left/ {\vphantom {{I_{1} } {K_{1} }}} \right. \kern-\nulldelimiterspace} {K_{1} }} \right)} \times {\sum\nolimits_{i = 1}^{n + 1} {{\left\lfloor {\prod ^{i}_{{j = 1}} K_{j} } \right\rfloor }} } \).
Therefore:
Any spectroscopic variable (absorbance, circular dichroism or fluorescence) at given wavelength can be expressed as:
Now using Eqs. (4) and (5), Eq. (6) can be rewritten as:
Using (from linear energy-model assumption):
Using the following approximation:
Equations (8) and (10) are used in Eqs. (1) and (2) of Materials and method. An intermediate I i is said to be spectroscopically silent at a particular wavelength λ, when \( <!!!>{\left( {\varepsilon ^{\lambda }_{N} - \varepsilon ^{\lambda }_{{I_{i} }} } \right)} = 0 \). In this case, though the corresponding components in Eqs. (8) and (10) become zero, the total free energy is an invariant, i.e., the free-energy component of the ith intermediate will be added to the free energy of the (i+1)th intermediate thus making the total free energy a constant.
Rights and permissions
About this article
Cite this article
Murugan, R., Mazumdar, S. Role of substrate on the conformational stability of the heme active site of cytochrome P450cam: effect of temperature and low concentrations of denaturants. J Biol Inorg Chem 9, 477–488 (2004). https://doi.org/10.1007/s00775-004-0544-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-004-0544-1