Abstract
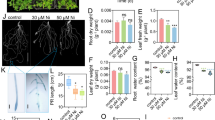
The present study is aimed at understanding the physiological and molecular responses of a wheat genotype (K9107), induced by the presence of P. minor in its vicinity, on root system architecture (RSA), 15N uptake, and the gene expression of TaNPF6.4 in seminal and lateral roots under optimal and NO3−-limiting conditions. Seedlings of both species were grown in a vermiculite-perlite mixture for a defined period until they achieved comparable biomass at the earliest. Then 2D images of roots were analyzed. The Kjeldahl and Isotope Ratio Mass Spectroscopy methods were used to measure total N and δ15N in wheat tissues. The transcript expression of all homeologs of the TaNPF6.4 gene was studied by Real-Time-Polymerase Chain Reaction (qPCR) analysis. The upstream sequences of TaNPF6.4 were cloned in the pJET1.2 cloning vector, and Cis-Regulatory Elements were identified using the PLACE database. Root length and fresh and dry weight of roots invariably increased under N-limiting conditions, which was further increased when grown in the vicinity of P. minor. Total root size (TRS), main root path length (MRP), lateral root size (LRS), and first and second-order lateral root numbers (FOLRN and SOLRN) of wheat seedlings increased when they grew in the vicinity of P. minor, especially under the N-limiting (0.08 mM NO3−) condition. The wheat seedling had the lowest 15N-enrichment in P. minor association under NO3−-limiting condition than other conditions. We identified four closest sequences orthologous to AtNRT1.1, a dual affinity nitrate transceptor, located on chromosomes 4, 5, and 7 of three sub-genomes of wheat, known as TaNPF6.4. The homeolog-specific expression of TaNPF6.4 revealed that the gene encoded by sub-genome A is the highest expressing homeolog in both seminal and lateral roots with enhanced relative expression under N-limiting conditions when grown along with P. minor. The upstream Cis-Regulatory Elements of all these four homeologous genes showed considerable variation in numbers. Our study demonstrates the modulation in architecture and development of wheat roots, differential expression of TaNPF6.4, and adverse effect on N uptake in P. minor association more severely under nitrate-limiting conditions, indicating a possible role of TaNPF6.4 in perceiving nitrate under kin and non-kin association.







Similar content being viewed by others
References
Badri DV, Vivanco JM (2009) Regulation and function of root exudates. Plant Cell Environ 32(6):666–681. https://doi.org/10.1111/j.1365-3040.2009.01926.x
Bhatt MV, Khandelwal A, Dudley SA (2011) Kin recognition, not competitive interactions, predicts root allocation in young Cakile edentula seedling pairs. New Phytol 189:1135–1142. https://doi.org/10.1111/j.1469-8137.2010.03548.x
Biedrzycki ML, Bais HP (2010) Kin recognition in plants: a mysterious behaviour unsolved. J Exp Bot 61:4123–4128. https://doi.org/10.1093/jxb/erq250
Bilas RD, Bretman A, Bennett T (2021) Friends, neighbours and enemies: an overview of the communal and social biology of plants. Plant Cell Environ 44:997–1013. https://doi.org/10.1111/pce.13965
Bouguyon E, Perrine-Walker F, Pervent M et al (2016) Nitrate controls root development through posttranscriptional regulation of the NRT1.1/NPF6.3 transporter/sensor. Plant Physiol 172:1237–1248. https://doi.org/10.1104/pp.16.01047
Buchner P, Hawkesford MJ (2014) Complex phylogeny and gene expression patterns of members of the nitrate transporter 1/peptide transporter family (NPF) in wheat. J Exp Bot 65:5697–5710. https://doi.org/10.1093/jxb/eru231
Chauhan BS (2020) Grand challenges in weed management. Front Agron 1:3. https://doi.org/10.3389/fagro.2019.00003
Chen BJ, During HJ, Anten NP (2012) Detect the neighbour: identity recognition at the root level in plants. Plant Sci 195:157–167. https://doi.org/10.1016/j.plantsci.2012.07.006
Chhokar RS, Malik RK (2002) Isoproturon resistant Phalaris minor and its response to alternate herbicides. Weed Technol 16:116–123
Chhokar RS, Sharma RK, Sharma I (2013) Weed management strategies in wheat-a review. J Wheat Res 4:1–21
Crepy MA, Casal JJ (2015) Photoreceptor mediated kin recognition in plants. New Phytol 205:329–338. https://doi.org/10.1111/nph.13040
Dudley SA, File AL (2007) Kin recognition in an annual plant. Biol Lett 3:435–438. https://doi.org/10.1098/rsbl.2007.0232
Garnett T, Conn V, Kaiser BN (2009) Root based approaches to improving nitrogen use efficiency in plants. Plant Cell Environ 32:1272–1283. https://doi.org/10.1111/j.1365-3040.2009.02011.x
Giehl RF, von Wiren N (2014) Root nutrient foraging. Plant Physiol 166:509–517. https://doi.org/10.1104/pp.114.245225
Giehl RF, Gruber BD, von Wiren N (2014) It’s time to make changes: modulation of root system architecture by nutrient signals. J Exp Bot 65:769–778. https://doi.org/10.1093/jxb/ert421
Gruber BD, Giehl RFH, Friedel S, Wiren N (2013) Plasticity of the Arabidopsis root system under nutrient deficiencies. Plant Physiol 168:161–179. https://doi.org/10.1104/pp.113.218453
Gruntman M, Novoplansky A (2004) Physiologically mediated self/non-self discrimination in roots. Proc Nat Acad Sci USA 101:3863–3867. https://doi.org/10.1073/pnas.0306604101
Harrington LW et al (1993) Wheat and rice in Karnal and Kurukshetra districts, Haryana, India: farmers’ practices, problems, and an agenda for action. D.F.; Haryana Agricultural University, Indian Council of Agricultural Research, International Maize and Wheat Improvement Center, and International Rice Research Institute, Mexico
Higo K, Ugawa Y, Iwamoto M (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300. https://doi.org/10.1093/nar/27.1.297
Ho CH, Tsay YF (2010) Nitrate, ammonium, and potassium sensing and signaling. Curr Opin Plant Biol 13:604–610. https://doi.org/10.1016/j.pbi.2010.08.005
Ho CH, Lin SH, Hu HC, Tsay YF (2009) CHL1 functions as a nitrate sensor in plants. Cell 138:1184–1194. https://doi.org/10.1016/j.cell.2009.07.004
Hofmann K, Bucher P, Falquet L, Bairoch A (1999) The PROSITE database, its status in 1999. Nucleic Acids Res 27:215–219. https://doi.org/10.1093/nar/27.1.215
Hu B, Wang W, Ou S, Tang J, Li H, Che R, Zhang Z, Chai X, Wang Y, Liang C (2015) Variation in NRT1.1B contributes to nitrate-use divergence between rice subspecies. Nat Genet 47:834. https://doi.org/10.1038/ng.3337
Huang NC, Chiang CS, Crawford NM, Tsay YF (1996) CHL1 encodes a component of the low-affinity nitrate uptake system in Arabidopsis and shows cell type-specific expression in roots. Plant Cell 8:2183–2191. https://doi.org/10.1105/tpc.8.12.2183
Hudson D, Guevara D, Yaish MW, Hannam C, Long N, Clarke JD, Bi YM, Rothstein SJ (2011) GNC and CGA1 modulate chlorophyll biosynthesis and glutamate synthase (GLU1/Fd-GOGAT) expression in Arabidopsis. PloS One 6:e26765. https://doi.org/10.1371/journal.pone.0026765
Imamura S, Kanesaki Y, Ohnuma M, Inouye T, Sekine Y, Fujiwara T, Kuroiwa T, Tanaka K (2009) R2R3-type MYB transcription factor, CmMYB1, is a central nitrogen assimilation regulator in Cyanidioschyzon merolae. Proc Nat Acad Sci USA 106:12548–12553. https://doi.org/10.1073/pnas.0902790106
Khera KL, Sandhu BS, Aujla TS, Singh CB, Kumar K (1995) Performance of wheat (Triticum aestivum) in relation to small canary grass (Phalaris minor) under different levels of irrigation, nitrogen and weed population. Indian J Agric Sci (India) 65:717–722 (https://epubs.icar.org.in/index.php/IJAgS/article/view/19085)
Krouk G, Crawford NM, Coruzzi GM, Tsay YF (2010) Nitrate signaling: adaptation to fluctuating environments. Curr Opin Plant Biol 13:265–272. https://doi.org/10.1016/j.pbi.2009.12.003
Krouk G, Lacombe B, Bielach A, Perrine-Walker F, Malinska K, Mounier E, Hoyerova K, Tillard P, Leon S, Ljung K, Zazimalova E, Benkova E, Nacry P, Gojon A (2010) Nitrate-regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants. Dev Cell 18:927–937. https://doi.org/10.1016/j.devcel.2010.05.008
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kumar A, Kumar S, Venkatesh K, Singh NK, Mandal PK, Sinha SK (2022) Physio-molecular traits of contrasting bread wheat genotypes associated with 15N influx exhibiting homeolog expression bias in nitrate transporter genes under different external nitrate concentrations. Planta 255:1–8. https://doi.org/10.1007/s00425-022-03890-7
Kumar A, Shamnas M, Jayaswal P, Mahato AK, Arya A, Mandal PK, Singh NK, Sinha SK (2023) Growth stage and nitrate limiting response of NRT2 and NAR2 gene families of bread wheat, and complementation and retrieval of nitrate uptake of antnrt2.1 mutant by a wheat NRT2 gene. Environ Exp Bot 207:105205. https://doi.org/10.1016/j.envexpbot.2022.105205
Ladha JK, Fischer KS, Hossain M, Hobbs PR, Hardy B (2000) Improving the productivity and sustainability of rice-wheat systems of the Indo-Gangetic plains: a synthesis of NARS-IRRI partnership research. Discussion Paper No. 40. International Rice Research Institute, Los Baños, Philippines. https://doi.org/10.22004/ag.econ.287597
Léran S, Muños S, Brachet C, Tillard P, Gojon A, Lacombe B (2013) Arabidopsis NRT1.1 is a bidirectional transporter involved in root-to-shoot nitrate translocation. Mol Plant 6:1984–1987. https://doi.org/10.1093/mp/sst068
Leran S, Varala K, Boyer JC, Chiurazzi M, Crawford N, Daniel-Vedele F, Gassmann W et al (2014) A unified nomenclature of nitrate transporter 1/peptide transporter family members in plants. Trends Plant Sci 9:5–9. https://doi.org/10.1016/j.tplants.2013.08.008
Li J, Xu X, Feng R (2018) Soil fertility and heavy metal pollution (Pb and Cd) alter kin interaction of Sorghum vulgare. Environ Exp Bot 155:368–377. https://doi.org/10.1016/j.envexpbot.2018.05.009
Li M, Tian H, Gao Y (2021) A genome-wide analysis of NPF and NRT2 transporter gene families in bread wheat provides new insights into the distribution, function, regulation and evolution of nitrate transporters. Plant Soil 465:47–63. https://doi.org/10.1007/s11104-021-04927-8
Liu Q, Wu K, Song W, Zhong N, Wu Y, Fu X (2022) Improving crop nitrogen use efficiency toward sustainable green revolution. Ann Rev Plant Biol 73:523–551. https://doi.org/10.1146/annurev-arplant-070121-015752
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2− ΔΔCT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Mahajan C, Sardana V (2013) Nutrient uptake by wheat and Phalaris minor as influenced by weed management practices. Agric Sci Digest 23:195–198
Mariotti A (1984) Natural 15N abundance measurements and atmospheric nitrogen standard calibration. Nature 311:251–252. https://doi.org/10.1038/311251a0
Medici A, Krouk G (2014) The primary nitrate response: a multifaceted signalling pathway. J Exp Bot 65:5567–5576. https://doi.org/10.1093/jxb/eru245
Melino VJ, Fiene G, Enju A, Cai J, Buchner P, Heuer S (2015) Genetic diversity for root plasticity and nitrogen uptake in wheat seedlings. Funct Plant Biol 42:942–956. https://doi.org/10.1071/FP15041
Mounier E, Pervent M, Ljung K, Gojon A, Nacry P (2014) Auxin-mediated nitrate signalling by NRT 1.1 participates in the adaptive response of Arabidopsis root architecture to the spatial heterogeneity of nitrate availability. Plant Cell Environ 37:162–174. https://doi.org/10.1111/pce.12143
Mukhopadhyay SK, Bera PS (1980) Effectiveness of basalin, tribunil, isoproturon and cultural methods in controlling weeds in wheat. Crop Pesticides 14:21–23
Orsel M, Krapp A, Daniel-Vedele F (2002) Analysis of the NRT2 nitrate transporter family in Arabidopsis. Structure and gene expression. Plant Physiol 129:886–896. https://doi.org/10.1104/pp.005280
Pezzola E, Pandolfi C, Mancuso S (2020) Resource availability affects kin selection in two cultivars of Pisum sativum. Plant Growth Reg 90:321–329. https://doi.org/10.1007/s10725-019-00562-7
Pierik R, Mommer L, Voesenek LACJ (2013) Molecular mechanisms of plant competition: neighbour detection and response strategies. Funct Ecol 27:841–853. https://doi.org/10.1111/1365-2435.12010
Ravishankar KV, Anand L, Dinesh MR (2000) Assessment of genetic relatedness among a few Indian mango cultivars using RAPD markers. J Hortic Sci Biotechnol 75:198–201. https://doi.org/10.1080/14620316.2000.11511223
Remans T, Nacry P, Pervent M, Filleur S, Diatloff E, Mounier E, Tillard P, Forde BG, Gojon A (2006) The Arabidopsis NRT1.1 transporter participates in the signaling pathway triggering root colonization of nitrate-rich patches. Proc Nat Acad Sci USA 103:19206–19211. https://doi.org/10.1073/pnas.060527510
Robinson D (1994) The responses of plants to non-uniform supplies of nutrients. New Phytol 127:635–674. https://doi.org/10.1111/j.1469-8137.1994.tb02969.x
Schenk HJ, Callaway RM, Mahall BE (1999) Spatial root segregation: are plants territorial? Adv Ecol Res 28:145–180. https://doi.org/10.1016/S0065-2504(08)60032-X
Sinha SK, Rani M, Bansal N, Venkatesh K, Mandal PK (2015) Nitrate starvation induced changes in root system architecture, carbon: nitrogen metabolism, and miRNA expression in nitrogen-responsive wheat genotypes. Appl Biochem Biotechnol 177:1299–1312. https://doi.org/10.1007/S12010-015-1815-8
Sinha SK, Rani M, Kumar A, Kumar S, Venkatesh K, Mandal PK (2018) Natural variation in root system architecture in diverse wheat genotypes grown under different nitrate conditions and root growth media. Theor Exp Plant Physiol 30:223–234. https://doi.org/10.1007/s40626-018-0117-2
Sinha SK, Tyagi A, Mandal PK (2019) External nitrogen and carbon source-mediated response on modulation of root system architecture and nitrate uptake in wheat seedlings. J Plant Growth Regul 38:283–297. https://doi.org/10.1007/s00344-018-9840-9
Sinha SK, Kumar A, Tyagi A, Venkatesh K, Paul D, Singh NK, Mandal PK (2020) Root architecture traits variation and nitrate-influx responses in diverse wheat genotypes under different external nitrogen concentrations. Plant Physiol Biochem 148:246–259. https://doi.org/10.1016/j.plaphy.2020.01.018
Timsina J, Connor DJ (2001) Productivity and management of rice-wheat cropping systems: issues and challenges. Field Crop Res 69:93–132. https://doi.org/10.1016/S0378-4290(00)00143-X
Tsay YF, Schroeder JI, Feldmann KA, Crawford NM (1993) The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 72:705–713. https://doi.org/10.1016/0092-8674(93)90399-b
Wang DJ, Liu Q, Lin JH, Sun RJ (2004) Optimum nitrogen use and reduced nitrogen loss for production of rice and wheat in the Yangtse Delta region. Environ Geochem Health 26:221–227. https://doi.org/10.1023/b:egah.0000039584.35434.e0
Wang H, Wan Y, Buchner P, King R, Ma H, Hawkesford MJ (2020) Phylogeny and gene expression of the complete nitrate transporter 1/peptide transporter family in Triticum aestivum. J Exp Bot 71:4531–4546. https://doi.org/10.1093/jxb/eraa210
Wang NQ, Kong CH, Wang P, Meiners SJ (2021) Root exudate signals in plant-plant interactions. Plant Cell Environ 44:1044–1058. https://doi.org/10.1111/pce.13892
Xu Y, Cheng HF, Kong CH, Meiners SJ (2021) Intra-specific kin recognition contributes to inter-specific allelopathy: a case study of allelopathic rice interference with paddy weeds. Plant Cell Environ 44:3709–3721. https://doi.org/10.1111/pce.14083
Yanagisawa S, Akiyama A, Kisaka H, Uchimiya H, Miwa T (2004) Metabolic engineering with Dof1 transcription factor in plants: improved nitrogen assimilation and growth under low-nitrogen conditions. Proc Nat Acad Sci USA 101:7833–7838. https://doi.org/10.1073/pnas.0402267101
Yang XF, Li LL, Xu Y, Kong CH (2018) Kin recognition in rice (Oryza sativa) lines. New Phytol 220:567–578. https://doi.org/10.1111/nph.15296
Zhang H, Forde BG (1998) An Arabidopsis MADS box gene that controls nutrient-induced changes in root architecture. Science 279:407–409. https://doi.org/10.1126/science.279.5349.407
Zhang L, Liu Q, Tian Y, Xu X, Ouyang H (2016) Kin selection or resource partitioning for growing with siblings: implications from measurements of nitrogen uptake. Plant Soil 398:79–86. https://doi.org/10.1007/s11104-015-2641-z
Funding
PK acknowledges financial support in the form of JRF to the Indian Council of Agricultural Research, New Delhi, India. The present study was financially supported by the Science & Engineering Research Board, Department of Science & Technology, Govt. of India, under the Core Research Grant (Plant Science; CRG/2018/00720) scheme and NIPB institutional in-house project on N-uptake. SKS is thankful to Dr. Subhash Chander and the Director, ICAR-Directorate of Weed Research, Jabalpur-482004, India, for providing seeds for Phalaris minor.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1:
Table S1 List of NRT1.1 gene identified in rice. Table S2 List of NRT1.1 gene identified in Wheat. Table S3 Homeologue specific primer sequence used for quantitative real time PCR. Table S4 Primer sequence for homeolog specific PCR amplification of upstream sequence. Fig. S1 Phylogenetic tree of NRT1.1 genes of rice and wheat. OsNRT1.1B (rice nitrate transceptor) and all wheat sequences orthologous to AtNRT1.1 (Arabidopsis nitrate transceptor) were used for construction phylogenetic tree.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kumari, P., Kumar, A., Kumar, S. et al. The Physio-molecular Responses of Bread Wheat (Triticum aestivum L.) Induced by Coexisting Phalaris minor Retz. Under Nitrate-limiting Conditions: Perspectives of Kin and Non-kin Recognition. J Soil Sci Plant Nutr 23, 3640–3654 (2023). https://doi.org/10.1007/s42729-023-01283-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-023-01283-7