Abstract
Background
We investigated whether the expression of transforming growth factor-beta-induced protein (TGFBI) and intratumoral immune cells including CD8- and Forkhead box protein P3 (Foxp3)-positive T cells in clinical lung cancer patients could predict the therapeutic response to nivolumab.
Methods
Thirty-three patients who were treated with nivolumab were enrolled in this study. Immunohistochemical analyses of TGFBI, PD-L1, CD8, Foxp3, and vimentin expression were conducted. Serum concentrations of TGFBI and transforming growth factor-beta1 (TGF-β1) were determined by enzyme-linked immunosorbent assay (ELISA).
Results
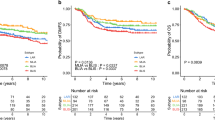
Cancer TGFBI was not associated with prognosis and therapeutic response to nivolumab, but cancer stromal TGFBI and intratumoral CD8-positive T cells were associated with them. Therefore, we evaluated cancer stromal TGFBI and intratumoral CD8-positive T cells. The high-TGFBI-expression group had poorer clinical responses than did the low-TGFBI-expression group (p < 0.0001). The number of times nivolumab was administered in the high-CD8-expression group was significantly higher than that in the low-CD8-expression group (p = 0.0046). The high-CD8-expression group had better clinical responses than did the low-CD8-expression group (p = 0.0013). Interestingly, all patients in the high-TGFBI/low-CD8-expression group had progressive disease (PD). In contrast, all patients in the low-TGFBI/high-CD8-expression group had PR + SD (partial response + stable disease) by the Response Evaluation Criteria in Solid Tumors (RECIST 1.1).
Conclusions
The dual evaluation of stromal TGFBI and intratumoral CD8-positive T cells could be a useful predictive marker for nivolumab.


Similar content being viewed by others
References
Youlden DR, Cramb SM, Baade PD. The international epidemiology of lung cancer: geographical distribution and secular trends. J Thorac Oncol. 2008;3(8):819–31.
Jemal A, Siegel R, Ward E, Hao Y, Xu J, Murray T, et al. Cancer statistics 2008. CA Cancer J Clin. 2008;58(2):71–96.
Okazaki T, Honjo T. The PD-1-PD-L pathway in immunological tolerance. Trends Immunol. 2006;27(4):195–201.
Keir ME, Liang SC, Guleria I, et al. Tissue expression of PD-L1 mediates peripheral T cell tolerance. J Exp Med. 2006;203(4):883–95.
Tseng SY, Otsuji M, Gorski K, et al. B7-DC, a new dendritic cell molecule with potent costimulatory properties for T cells. J Exp Med. 2001;193(7):839–46.
Dong H, Strome SE, Salomao DR, et al. Tumor-associated B7-H1 promotes T-cell apoptosis: a potential mechanism of immune evasion. Nat Med. 2002;8(8):793–800.
Iwai Y, Ishida M, Tanaka Y, Okazaki T, Honjo T, Minato N. Involvement of PD-L1 on tumor cells in the escape from host immune system and tumor immunotherapy by PD-L1 blockade. Proc Natl Acad Sci USA. 2002;99(19):12293–7.
Tsushima F, Yao S, Shin T, et al. Interaction between B7-H1 and PD-1 determines initiation and reversal of T-cell anergy. Blood. 2007;110(1):180–5.
Brahmer J, Reckamp KL, Baas P, et al. Nivolumab versus docetaxel in advanced squamous-cell non-small-cell lung cancer. N Engl J Med. 2015;373(2):123–35.
Borghaei H, Paz-Ares L, Horn L, et al. Nivolumab versus docetaxel in advanced nonsquamous non-small-cell lung cancer. N Engl J Med. 2015;373(17):1627–39.
Reck M, Rodríguez-Abreu D, Robinson AG, et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N Engl J Med. 2016;375(19):1823–33.
Dempke WCM, Fenchel K, Dale SP. Programmed cell death ligand-1 (PD-L1) as a biomarker for non-small cell lung cancer (NSCLC) treatment-are we barking up the wrong tree? Transl Lung Cancer Res. 2018;7(Suppl 3):275–9.
Lim SH, Hong M, Ahn S, et al. Changes in tumour expression of programmed death-ligand 1 after neoadjuvant concurrent chemoradiotherapy in patients with squamous oesophageal cancer. Eur J Cancer. 2016;52:1–9.
McLaughlin J, Han G, Schalper KA, et al. Quantitative assessment of the heterogeneity of PD-L1 expression in non-small-cell lung cancer. JAMA Oncol. 2016;2(1):46–54.
Peters S, Creelan B, Hellmann MD, et al. Abstract CT082: impact of tumor mutation burden on the efficacy of first-line nivolumab in stage iv or recurrent non-small cell lung cancer: an exploratory analysis of CheckMate 026. Cancer Res. 2017;77(Supplement 13):CT082.
Mariathasan S, Turley SJ, Nickles D, et al. TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells. Nature. 2018;554(7693):544–8.
Yin J, Lu K, Lin J, et al. Genetic variants in TGF-β pathway are associated with ovarian cancer risk. PLoS ONE. 2011;6(9):e25559.
Scollen S, Luccarini C, Baynes C, et al. TGF-β signaling pathway and breast cancer susceptibility. Cancer Epidemiol Biomarkers Prev. 2011;20(6):1112–9.
Fang F, Yu L, Zhong Y, Yao L. TGFB1 509 C/T polymorphism and colorectal cancer risk: a meta-analysis. Med Oncol. 2010;27(4):1324–8.
Bhayal AC, Prabhakar B, Rao KP, et al. Role of transforming growth factor-β1 -509 C/T promoter polymorphism in gastric cancer in south Indian population. Tumour Biol. 2011;32(5):1049–53.
Lin M, Stewart DJ, Spitz MR, et al. Genetic variations in the transforming growth factor-beta pathway as predictors of survival in advanced non-small cell lung cancer. Carcinogenesis. 2011;32(7):1050–6.
Javle M, Li Y, Tan D, Dong X, Chang P, Kar S, et al. Biomarkers of TGF-β signaling pathway and prognosis of pancreatic cancer. PLoS ONE. 2014;9(1):e85942.
Costanza B, Umelo IA, Bellier J, Castronovo V, Turtoi A. Stromal modulators of TGF-β in cancer. J Clin Med. 2017;6(1):E7.
Skonier J, Neubauer M, Madisen L, Bennett K, Plowman GD, Purchio AF. cDNA cloning and sequence analysis of βig-h3, a novel gene induced in a human adenocarcinoma cell line after treatment with transforming growth factor-beta. DNA Cell Biol. 1992;11(7):511–22.
Turtoi A, Musmeci D, Wang Y, et al. Identification of novel accessible proteins bearing diagnostic and therapeutic potential in human pancreatic ductal adenocarcinoma. J Proteome Res. 2011;10(9):4302–13.
Turtoi A, Blomme A, Debois D, et al. Organized proteomic heterogeneity in colorectal cancer liver metastases and implications for therapies. Hepatology. 2014;59(3):924–34.
Kawamoto T, Noshiro M, Shen M, et al. Structural and phylogenetic analyses of RGD-CAP/βig-h3, a fasciclin-like adhesion protein expressed in chick chondrocytes. Biochim Biophys Acta. 1998;1395(3):288–92.
Yokobori T, Nishiyama M. TGF-β signaling in gastrointestinal cancers: progress in basic and clinical research. J Clin Med. 2017;6(1):E11.
Liu Y, Xue M, Du S, et al. Competitive endogenous RNA is an intrinsic component of EMT regulatory circuits and modulates EMT. Nat Commun. 2019;10(1):1637.
Wang J, Chen Y, Xiang F, et al. Suppression of TGF-β1 enhances chemosensitivity of cisplatin-resistant lung cancer cells through the inhibition of drug-resistant proteins. Artif Cells Nanomed Biotechnol. 2018;46(7):1505–12.
Kaira K, Higuchi T, Naruse I, et al. Metabolic activity by 18F-FDG-PET/CT is predictive of early response after nivolumab in previously treated NSCLC. Eur J Nucl Med Mol Imaging. 2018;45(1):56–66.
Shimoda Y, Ubukata Y, Handa T, et al. High expression of forkhead box protein C2 is associated with aggressive phenotypes and poor prognosis in clinical hepatocellular carcinoma. BMC Cancer. 2018;18(1):597.
Fong YC, Hsu SF, Wu CL, et al. Transforming growth factor-beta1 increases cell migration and beta1 integrin up-regulation in human lung cancer cells. Lung Cancer. 2009;64(1):13–21.
Sasaki H, Kobayashi Y, Nakashima Y, et al. Beta IGH3, a TGF-beta inducible gene, is overexpressed in lung cancer. Jpn J Clin Oncol. 2002;32(3):85–9.
O’Leary K, Shia A, Cavicchioli F, et al. Identification of Endoglin as an epigenetically regulated tumour-suppressor gene in lung cancer. Br J Cancer. 2015;113(6):970–8.
Wu X, Ruan L, Yang Y, Mei Q. Analysis of gene expression changes associated with human carcinoma-associated fibroblasts in non-small cell lung carcinoma. Biol Res. 2017;50(1):6.
Kim JE, Kim SJ, Lee BH, Park RW, Kim KS, Kim IS. Identification of motifs for cell adhesion within the repeated domains of transforming growth factor-beta-induced gene, betaig-h3. J Biol Chem. 2000;275(40):30907–15.
Bhagirath D, Abrol N, Khan R, Sharma M, Seth A, Sharma A. Expression of CD147, BIGH3 and Stathmin and their potential role as diagnostic marker in patients with urothelial carcinoma of the bladder. Clin Chim Acta. 2012;413(19–20):1641–6.
Han B, Cai H, Chen Y, Hu B, Luo H, Wu Y, et al. The role of TGFBI (βig-H3) in gastrointestinal tract tumorigenesis. Mol Cancer. 2015;14:64.
Mu CY, Huang JA, Chen Y, Chen C, Zhang XG. High expression of PD-L1 in lung cancer may contribute to poor prognosis and tumor cells immune escape through suppressing tumor infiltrating dendritic cells maturation. Med Oncol. 2011;28(3):682–8.
Konishi J, Yamazaki K, Azuma M, Kinoshita I, Dosaka-Akita H, Nishimura M. B7-H1 expression on non-small cell lung cancer cells and its relationship with tumor-infiltrating lymphocytes and their PD-1 expression. Clin Cancer Res. 2004;10(15):5094–100.
El-Guindy DM, Helal DS, Sabry NM, Abo El-Nasr M. Programmed cell death ligand-1 (PD-L1) expression combined with CD8 tumor infiltrating lymphocytes density in non-small cell lung cancer patients. J Egypt Natl Canc Inst. 2018;30(4):125–31.
Tumeh PC, Harview CL, Yearley JH, et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature. 2014;515(7528):568–71.
Zou W, Wolchok JD, Chen L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci Transl Med. 2016;8(328):328rv4.
Lanitis E, Dangaj D, Irving M, Coukos G. Mechanisms regulating T-cell infiltration and activity in solid tumors. Ann Oncol. 2017;28:xii18–32.
Wang L, Saci A, Szabo PM, et al. EMT-and stroma-related gene expression and resistance to PD-1 blockade in urothelial cancer. Nat Commun. 2018;9(1):3503.
Zhang L, Chen Y, Li F, Bao L, Liu W. Atezolizumab and bevacizumab attenuate cisplatin resistant ovarian cancer cells progression synergistically via suppressing epithelial-mesenchymal transition. Front Immunol. 2019;10:867.
Chae YK, Chang S, Ko T, et al. Epithelial-mesenchymal transition (EMT) signature is inversely associated with T-cell infiltration in non-small cell lung cancer (NSCLC). Sci Rep. 2018;8(1):2918.
Sato H, Niimi A, Yasuhara T, et al. DNA double-strand break repair pathway regulates PD-L1 expression in cancer cells. Nat Commun. 2017;8(1):1751.
Acknowledgment
The authors thank Ms. Yukie Saito, Ms. Sayaka Okada, Ms. Kayoko Takahashi, Ms. Mizue Murata, Ms. Harumi Kanai, Ms. Fumie Takada, Ms. Sawa Nagayama, and Ms. Mariko Nakamura for their excellent assistance. This work was supported by the research grant from Ono Pharmaceutical Co., Ltd. and Bristol-Myers Squibb K.K and Grants-in-Aid for Scientific Research from the Japan Society for the Promotion of Science (JSPS), grant numbers 17K19893, 18K07665, and 18H02877. The work also was supported in part by a Research Grant of the Princess Takamatsu Cancer Research Fund, the Suzuken Memorial Foundation, and the Pancreas Research Foundation of Japan.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Disclosures
Kyoichi Kaira has received research grants and a speaker honorarium from ONO PHARMACEUTICAL CO., LTD. and Bristol-Myers Squibb K.K.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nakazawa, N., Yokobori, T., Kaira, K. et al. High Stromal TGFBI in Lung Cancer and Intratumoral CD8-Positive T Cells were Associated with Poor Prognosis and Therapeutic Resistance to Immune Checkpoint Inhibitors. Ann Surg Oncol 27, 933–942 (2020). https://doi.org/10.1245/s10434-019-07878-8
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-019-07878-8