Abstract
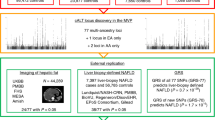
Nonalcoholic fatty liver disease (NAFLD) is common and partially heritable and has no effective treatments. We carried out a genome-wide association study (GWAS) meta-analysis of imaging (n = 66,814) and diagnostic code (3,584 cases versus 621,081 controls) measured NAFLD across diverse ancestries. We identified NAFLD-associated variants at torsin family 1 member B (TOR1B), fat mass and obesity associated (FTO), cordon-bleu WH2 repeat protein like 1 (COBLL1)/growth factor receptor-bound protein 14 (GRB14), insulin receptor (INSR), sterol regulatory element-binding transcription factor 1 (SREBF1) and patatin-like phospholipase domain-containing protein 2 (PNPLA2), as well as validated NAFLD-associated variants at patatin-like phospholipase domain-containing protein 3 (PNPLA3), transmembrane 6 superfamily 2 (TM6SF2), apolipoprotein E (APOE), glucokinase regulator (GCKR), tribbles homolog 1 (TRIB1), glycerol-3-phosphate acyltransferase (GPAM), mitochondrial amidoxime-reducing component 1 (MARC1), microsomal triglyceride transfer protein large subunit (MTTP), alcohol dehydrogenase 1B (ADH1B), transmembrane channel like 4 (TMC4)/membrane-bound O-acyltransferase domain containing 7 (MBOAT7) and receptor-type tyrosine-protein phosphatase δ (PTPRD). Implicated genes highlight mitochondrial, cholesterol and de novo lipogenesis as causally contributing to NAFLD predisposition. Phenome-wide association study (PheWAS) analyses suggest at least seven subtypes of NAFLD. Individuals in the top 10% and 1% of genetic risk have a 2.5-fold to 6-fold increased risk of NAFLD, cirrhosis and hepatocellular carcinoma. These genetic variants identify subtypes of NAFLD, improve estimates of disease risk and can guide the development of targeted therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Meta-analysis results from this study are available at http://www.med.umich.edu/spelioteslab/ and at GWAS Catalog (GCP ID: GCP000662). GOLD Consortium and MGI individual-level data are governed by patient privacy requirements and available to those having the mandatory IRB approvals. The eMERGE NAFLD cohort was previously described, and summary statistics are publicly available (https://www.ebi.ac.uk/gwas/studies/GCST008468). FinnGen data freeze 4 summary statistics are publicly available (https://www.finngen.fi/fi). UKBB genomic and phenotypic data supporting this publication are available upon application (https://ukbiobank.ac.uk). Otherwise, all data used to generate figures can be found in supplementary tables or in the above publicly available datasets. Source data are provided with this paper.
Code availability
Data analyses were performed using publicly available codes or software.
References
Lazo, M. et al. Prevalence of nonalcoholic fatty liver disease in the United States: the third National Health and Nutrition Examination Survey, 1988–1994. Am. J. Epidemiol. 178, 38–45 (2013).
Portillo Sanchez, P. et al. High prevalence of nonalcoholic fatty liver disease in patients with type 2 diabetes mellitus and normal plasma aminotransferase levels. J. Clin. Endocrinol. Metab. 100, 2231–2238 (2015).
Dongiovanni, P. et al. Causal relationship of hepatic fat with liver damage and insulin resistance in nonalcoholic fatty liver. J. Intern. Med. 283, 356–370 (2018).
Lauridsen, B. K. et al. Liver fat content, non-alcoholic fatty liver disease, and ischaemic heart disease: Mendelian randomization and meta-analysis of 279 013 individuals. Eur. Heart J. 39, 385–393 (2018).
Stender, S. et al. Adiposity amplifies the genetic risk of fatty liver disease conferred by multiple loci. Nat. Genet. 49, 842–847 (2017).
Liu, Z. et al. Causal relationships between NAFLD, T2D and obesity have implications for disease subphenotyping. J. Hepatol. 73, 263–276 (2020).
Bianco, C. et al. Non-invasive stratification of hepatocellular carcinoma risk in non-alcoholic fatty liver using polygenic risk scores. J. Hepatol. 74, 775–782 (2021).
Parisinos, C. A. et al. Genome-wide and Mendelian randomisation studies of liver MRI yield insights into the pathogenesis of steatohepatitis. J. Hepatol. 73, 241–251 (2020).
Crespo, J. et al. Are there predictive factors of severe liver fibrosis in morbidly obese patients with non-alcoholic steatohepatitis? Obes. Surg. 11, 254–257 (2001).
Younossi, Z. M. et al. The economic and clinical burden of nonalcoholic fatty liver disease in the United States and Europe. Hepatology 64, 1577–1586 (2016).
Romeo, S. et al. Genetic variation in PNPLA3 confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 40, 1461–1465 (2008).
Speliotes, E. K. et al. Genome-wide association analysis identifies variants associated with nonalcoholic fatty liver disease that have distinct effects on metabolic traits. PLoS Genet. 7, e1001324 (2011).
Luukkonen, P. K. et al. MARC1 variant rs2642438 increases hepatic phosphatidylcholines and decreases severity of non-alcoholic fatty liver disease in humans. J. Hepatol. 73, 725–726 (2020).
Jamialahmadi, O. et al. Exome-wide association study on alanine aminotransferase identifies sequence variants in the GPAM and APOE associated with fatty liver disease. Gastroenterology 160, 1634–1646 (2021).
Kitamoto, A. et al. Association of polymorphisms in GCKR and TRIB1 with nonalcoholic fatty liver disease and metabolic syndrome traits. Endocr. J. 61, 683–689 (2014).
Mancina, R. M. et al. The MBOAT7-TMC4 variant rs641738 increases risk of nonalcoholic fatty liver disease in individuals of European descent. Gastroenterology 150, 1219–1230 (2016).
Nakajima, S. et al. Polymorphism of receptor-type tyrosine-protein phosphatase δ gene in the development of non-alcoholic fatty liver disease. J. Gastroenterol. Hepatol. 33, 283–290 (2018).
Palmer, N. D. et al. Allele-specific variation at APOE increases nonalcoholic fatty liver disease and obesity but decreases risk of Alzheimer’s disease and myocardial infarction. Hum. Mol. Genet. 30, 1443–1456 (2021).
Vilar-Gomez, E. et al. ADH1B*2 is associated with reduced severity of nonalcoholic fatty liver disease in adults, independent of alcohol consumption. Gastroenterology 159, 929–943 (2020).
Zheng, W. et al. MTP -493G>T polymorphism and susceptibility to nonalcoholic fatty liver disease: a meta-analysis. DNA Cell Biol. 33, 361–369 (2014).
Middleton, M. S. et al. Agreement between magnetic resonance imaging proton density fat fraction measurements and pathologist-assigned steatosis grades of liver biopsies from adults with nonalcoholic steatohepatitis. Gastroenterology 153, 753–761 (2017).
Saadeh, S. et al. The utility of radiological imaging in nonalcoholic fatty liver disease. Gastroenterology 123, 745–750 (2002).
Pers, T. H. et al. Biological interpretation of genome-wide association studies using predicted gene functions. Nat. Commun. 6, 5890 (2015).
Kahali, B. et al. A noncoding variant near PPP1R3B promotes liver glycogen storage and MetS, but protects against myocardial infarction. J. Clin. Endocrinol. Metab. 106, 372–387 (2021).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Lawlor, D. A. et al. Mendelian randomization: using genes as instruments for making causal inferences in epidemiology. Stat. Med. 27, 1133–1163 (2008).
Chen, V. L. et al. Genome-wide association study of serum liver enzymes implicates diverse metabolic and liver pathology. Nat. Commun. 12, 816 (2021).
Emdin, C. A. et al. Association of genetic variation with cirrhosis: a multi-trait genome-wide association and gene-environment interaction study. Gastroenterology 160, 1620–1633 (2021).
Sveinbjornsson, G. et al. Multiomics study of nonalcoholic fatty liver disease. Nat. Genet. 54, 1652–1663 (2022).
Vujkovic, M. et al. A multiancestry genome-wide association study of unexplained chronic ALT elevation as a proxy for nonalcoholic fatty liver disease with histological and radiological validation. Nat. Genet. 54, 761–771 (2022).
Chambers, J. C. et al. Genome-wide association study identifies loci influencing concentrations of liver enzymes in plasma. Nat. Genet. 43, 1131–1138 (2011).
Kathiresan, S. et al. Six new loci associated with blood low-density lipoprotein cholesterol, high-density lipoprotein cholesterol or triglycerides in humans. Nat. Genet. 40, 189–197 (2008).
Beer, N. L. et al. The P446L variant in GCKR associated with fasting plasma glucose and triglyceride levels exerts its effect through increased glucokinase activity in liver. Hum. Mol. Genet. 18, 4081–4088 (2009).
Ishizuka, Y. et al. TRIB1 downregulates hepatic lipogenesis and glycogenesis via multiple molecular interactions. J. Mol. Endocrinol. 52, 145–158 (2014).
Bauer, R. C. et al. Tribbles-1 regulates hepatic lipogenesis through posttranscriptional regulation of C/EBPalpha. J. Clin. Invest. 125, 3809–3818 (2015).
Agius, L. Hormonal and metabolite regulation of hepatic glucokinase. Annu. Rev. Nutr. 36, 389–415 (2016).
Janssen, M. C. et al. Symptomatic lipid storage in carriers for the PNPLA2 gene. Eur. J. Hum. Genet. 21, 807–815 (2013).
Steneberg, P. et al. Hyperinsulinemia enhances hepatic expression of the fatty acid transporter Cd36 and provokes hepatosteatosis and hepatic insulin resistance. J. Biol. Chem. 290, 19034–19043 (2015).
Ipsen, D. H., Lykkesfeldt, J. & Tveden-Nyborg, P. Molecular mechanisms of hepatic lipid accumulation in non-alcoholic fatty liver disease. Cell. Mol. Life Sci. 75, 3313–3327 (2018).
Popineau, L. et al. Novel Grb14-mediated cross talk between insulin and p62/Nrf2 pathways regulates liver lipogenesis and selective insulin resistance. Mol. Cell. Biol. 36, 2168–2181 (2016).
Cooney, G. J. et al. Improved glucose homeostasis and enhanced insulin signalling in Grb14-deficient mice. EMBO J. 23, 582–593 (2004).
Michael, M. D. et al. Loss of insulin signaling in hepatocytes leads to severe insulin resistance and progressive hepatic dysfunction. Mol. Cell 6, 87–97 (2000).
Sirwi, A. & Hussain, M. M. Lipid transfer proteins in the assembly of apoB-containing lipoproteins. J. Lipid Res. 59, 1094–1102 (2018).
Adam, M. P. et al. (eds.). GeneReviews((R)) (University of Washington, 1993).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Polimanti, R. & Gelernter, J. ADH1B: from alcoholism, natural selection, and cancer to the human phenome. Am. J. Med. Genet. B Neuropsychiatr. Genet. 177, 113–125 (2018).
Gu, S. et al. Recent selection on a class I ADH locus distinguishes Southwest Asian populations including Ashkenazi Jews. Genes (Basel) 9, 452 (2018).
Macgregor, S. et al. Associations of ADH and ALDH2 gene variation with self report alcohol reactions, consumption and dependence: an integrated analysis. Hum. Mol. Genet. 18, 580–593 (2009).
Muenter, M. D., Perry, H. O. & Ludwig, J. Chronic vitamin A intoxication in adults. Hepatic, neurologic and dermatologic complications. Am. J. Med. 50, 129–136 (1971).
Shin, J. Y. et al. Nuclear envelope-localized torsinA-LAP1 complex regulates hepatic VLDL secretion and steatosis. J. Clin. Invest. 129, 4885–4900 (2019).
Innes, H. et al. Genome-wide association study for alcohol-related cirrhosis identifies risk loci in MARC1 and HNRNPUL1. Gastroenterology 159, 1276–1289.e7 (2020).
Xia, M. et al. Hepatic deletion of Mboat7 (Lpiat1) causes activation of SREBP-1c and fatty liver. J. Lipid Res. 62, 100031 (2021).
Landgraf, K. et al. FTO obesity risk variants are linked to adipocyte IRX3 expression and BMI of children—relevance of FTO variants to defend body weight in lean children? PLoS ONE 11, e0161739 (2016).
Kozlitina, J. et al. Exome-wide association study identifies a TM6SF2 variant that confers susceptibility to nonalcoholic fatty liver disease. Nat. Genet. 46, 352–356 (2014).
Wang, Y. et al. PNPLA3, CGI-58, and inhibition of hepatic triglyceride hydrolysis in mice. Hepatology 69, 2427–2441 (2019).
Morton, A. M. et al. Apolipoproteins E and CIII interact to regulate HDL metabolism and coronary heart disease risk. JCI Insight 3, e98045 (2018).
Abul-Husn, N. S. et al. A protein-truncating HSD17B13 variant and protection from chronic liver disease. N. Engl. J. Med. 378, 1096–1106 (2018).
Fox, C. S. et al. Genome-wide association for abdominal subcutaneous and visceral adipose reveals a novel locus for visceral fat in women. PLoS Genet. 8, e1002695 (2012).
Hatters, D. M., Peters-Libeu, C. A. & Weisgraber, K. H. Apolipoprotein E structure: insights into function. Trends Biochem. Sci. 31, 445–454 (2006).
Mahdessian, H. et al. TM6SF2 is a regulator of liver fat metabolism influencing triglyceride secretion and hepatic lipid droplet content. Proc. Natl Acad. Sci. USA 111, 8913–8918 (2014).
BasuRay, S. et al. Accumulation of PNPLA3 on lipid droplets is the basis of associated hepatic steatosis. Proc. Natl Acad. Sci. USA 116, 9521–9526 (2019).
Rondinone, C. M. et al. Protein tyrosine phosphatase 1B reduction regulates adiposity and expression of genes involved in lipogenesis. Diabetes 51, 2405–2411 (2002).
Zou, Y. et al. IRX3 promotes the browning of white adipocytes and its rare variants are associated with human obesity risk. EBioMedicine 24, 64–75 (2017).
Zeng, H. et al. CD36 promotes de novo lipogenesis in hepatocytes through INSIG2-dependent SREBP1 processing. Mol. Metab. 57, 101428 (2022).
Cignarelli, A. et al. Insulin and insulin receptors in adipose tissue development. Int. J. Mol. Sci. 20, 759 (2019).
Ong, K. T. et al. Adipose triglyceride lipase is a major hepatic lipase that regulates triacylglycerol turnover and fatty acid signaling and partitioning. Hepatology 53, 116–126 (2011).
Morales, L. D. et al. Further evidence supporting a potential role for ADH1B in obesity. Sci. Rep. 11, 1932 (2021).
Tanaka, Y. et al. LPIAT1/MBOAT7 depletion increases triglyceride synthesis fueled by high phosphatidylinositol turnover. Gut 70, 180–193 (2021).
Neschen, S. et al. Prevention of hepatic steatosis and hepatic insulin resistance in mitochondrial acyl-CoA:glycerol-sn-3-phosphate acyltransferase 1 knockout mice. Cell Metab. 2, 55–65 (2005).
Linden, D. et al. Liver-directed overexpression of mitochondrial glycerol-3-phosphate acyltransferase results in hepatic steatosis, increased triacylglycerol secretion and reduced fatty acid oxidation. FASEB J. 20, 434–443 (2006).
Klein, J. M. et al. The mitochondrial amidoxime-reducing component (mARC1) is a novel signal-anchored protein of the outer mitochondrial membrane. J. Biol. Chem. 287, 42795–42803 (2012).
Hussain, M. M. et al. Multiple functions of microsomal triglyceride transfer protein. Nutr. Metab. (Lond.) 9, 14 (2012).
Fernandes Silva, L. et al. An intronic variant in the GCKR gene is associated with multiple lipids. Sci. Rep. 9, 10240 (2019).
Douvris, A. et al. Functional analysis of the TRIB1 associated locus linked to plasma triglycerides and coronary artery disease. J. Am. Heart Assoc. 3, e000884 (2014).
Harris, T. B. et al. Age, Gene/Environment Susceptibility-Reykjavik Study: multidisciplinary applied phenomics. Am. J. Epidemiol. 165, 1076–1087 (2007).
Regan, E. A. et al. Genetic epidemiology of COPD (COPDGene) study design. COPD 7, 32–43 (2010).
Carr, J. J. et al. Calcified coronary artery plaque measurement with cardiac CT in population-based studies: standardized protocol of Multi-Ethnic Study of Atherosclerosis (MESA) and Coronary Artery Risk Development in Young Adults (CARDIA) study. Radiology 234, 35–43 (2005).
Speliotes, E. K. et al. Liver fat is reproducibly measured using computed tomography in the Framingham Heart Study. J. Gastroenterol. Hepatol. 23, 894–899 (2008).
Daniels, P. R. et al. Familial aggregation of hypertension treatment and control in the Genetic Epidemiology Network of Arteriopathy (GENOA) study. Am. J. Med. 116, 676–681 (2004).
Palmer, N. D. et al. Genetic variants associated with quantitative glucose homeostasis traits translate to type 2 diabetes in Mexican Americans: the GUARDIAN (Genetics Underlying Diabetes in Hispanics) Consortium. Diabetes 64, 1853–1866 (2015).
Liu, J. et al. Fatty liver, abdominal adipose tissue and atherosclerotic calcification in African Americans: the Jackson Heart Study. Atherosclerosis 224, 521–525 (2012).
Kramer, H. et al. Racial/ethnic differences in hypertension and hypertension treatment and control in the multi-ethnic study of atherosclerosis (MESA). Am. J. Hypertens. 17, 963–970 (2004).
Rampersaud, E. et al. The association of coronary artery calcification and carotid artery intima-media thickness with distinct, traditional coronary artery disease risk factors in asymptomatic adults. Am. J. Epidemiol. 168, 1016–1023 (2008).
Canela-Xandri, O., Rawlik, K. & Tenesa, A. An atlas of genetic associations in UK Biobank. Nat. Genet. 50, 1593–1599 (2018).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation (Springer International Publishing, 2015).
Yushkevich, P. A. et al. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage 31, 1116–1128 (2006).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proc. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 770–778 (IEEE, 2016).
Namjou, B. et al. GWAS and enrichment analyses of non-alcoholic fatty liver disease identify new trait-associated genes and pathways across eMERGE Network. BMC Med. 17, 135 (2019).
Chen, V. L. et al. Genetic variants that associate with cirrhosis have pleiotropic effects on human traits. Liver Int. 40, 405–415 (2020).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Zhou, W. et al. Efficiently controlling for case-control imbalance and sample relatedness in large-scale genetic association studies. Nat. Genet. 50, 1335–1341 (2018).
Dongiovanni, P. et al. Genetic variants regulating insulin receptor signalling are associated with the severity of liver damage in patients with non-alcoholic fatty liver disease. Gut 59, 267–273 (2010).
Feitosa, M. F. et al. The ERLIN1-CHUK-CWF19L1 gene cluster influences liver fat deposition and hepatic inflammation in the NHLBI Family Heart Study. Atherosclerosis 228, 175–180 (2013).
Chalasani, N. et al. Genome-wide association study identifies variants associated with histologic features of nonalcoholic fatty liver disease. Gastroenterology 139, 1567–1576 (2010).
Eslam, M. et al. Interferon-λ rs12979860 genotype and liver fibrosis in viral and non-viral chronic liver disease. Nat. Commun. 6, 6422 (2015).
Wiedmann, S. et al. Genetic variants within the LPIN1 gene, encoding lipin, are influencing phenotypes of the metabolic syndrome in humans. Diabetes 57, 209–217 (2008).
Shang, X. R. et al. GWAS-identified common variants with nonalcoholic fatty liver disease in Chinese children. J. Pediatr. Gastroenterol. Nutr. 60, 669–674 (2015).
Petta, S. et al. IL28B and PNPLA3 polymorphisms affect histological liver damage in patients with non-alcoholic fatty liver disease. J. Hepatol. 56, 1356–1362 (2012).
Kitamoto, T. et al. Genome-wide scan revealed that polymorphisms in the PNPLA3, SAMM50, and PARVB genes are associated with development and progression of nonalcoholic fatty liver disease in Japan. Hum. Genet. 132, 783–792 (2013).
Anstee, Q. M. et al. Genome-wide association study of non-alcoholic fatty liver and steatohepatitis in a histologically characterised cohort. J. Hepatol. 73, 505–515 (2020).
Ma, Y. et al. 17-β hydroxysteroid dehydrogenase 13 is a hepatic retinol dehydrogenase associated with histological features of nonalcoholic fatty liver disease. Hepatology 69, 1504–1519 (2019).
Park, S. L. et al. Genome-wide association study of liver fat: the Multiethnic Cohort Adiposity Phenotype Study. Hepatol. Commun. 4, 1112–1123 (2020).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Hemani, G. et al. The MR-base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Bowden, J. et al. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: the role of the I2 statistic. Int. J. Epidemiol. 45, 1961–1974 (2016).
Acknowledgements
AGES was funded by the National Institutes of Health (NIH; contracts N01-AG-1-2100 and HHSN271201200022C), the NIA Intramural Research Program, Hjartavernd (the Icelandic Heart Association) and the Althingi (the Icelandic Parliament). Support for FamHS was provided by the National Heart, Lung and Blood Institute (NHLBI; grants R01 HL087700 and R01 HL117078) and the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK; grant R01 DK089256 to M.A.P.). FHS is conducted and supported by the NHLBI in collaboration with Boston University (contracts N01-HC-25195, HHSN268201500001I and 75N92019D00031). Funding for SHARe Affymetrix genotyping was provided by NHLBI (contract N02-HL64278). SHARe Illumina genotyping was provided under an agreement between Illumina and Boston University. The Old Order Amish liver phenotyping is supported by NIH grants and contracts (U01 HL072515 and P30 DK72488) and analysis methods by U01 HL137181 (to J.R.O.). Support for the GENOA study was provided by the NIH (grants HL085571 to P.A.P. and HL087660 to S.L.R.K.) and NHLBI (HL100245). Support for the IRASFS was provided by the NHLBI (grants R01 HL060944, R01 HL061019, R01 HL060919, R01 HL060894 and R01 HL061210 to X.G., D.W.B., J.M.N., J.I.R., L.E.W. and N.D.P.). Genotyping and analysis were supported by NIDDK (grants DK085175 and R01 DK118062). JHS is supported and conducted in collaboration with Jackson State University (HHSN268201800013I), Tougaloo College (HHSN268201800014I), the Mississippi State Department of Health (HHSN268201800015I) and the University of Mississippi Medical Center (HHSN268201800010I, HHSN268201800011I and HHSN268201800012I) contracts from the NHLBI and the National Institute on Minority Health and Health Disparities (NIMHD). We also thank the staff and participants of the JHS. MESA and the MESA SHARe projects are conducted and supported by the NHLBI in collaboration with MESA investigators. Support for MESA is provided by contracts 75N92020D00001, HHSN268201500003I, N01-HC-95159, 75N92020D00005, N01-HC-95160, 75N92020D00002, N01-HC-95161, 75N92020D00003, N01-HC-95162, 75N92020D00006, N01-HC-95163, 75N92020D00004, N01-HC-95164, 75N92020D00007, N01-HC-95165, N01-HC-95166, N01-HC-95167, N01-HC-95168, N01-HC-95169, UL1-TR-000040, UL1-TR-001079 and UL1-TR-001420, UL1TR001881, DK063491 and R01HL105756. Funding for SHARe genotyping was provided by NHLBI (contract N02-HL-64278). Genotyping was performed at Affymetrix and the Broad Institute of Harvard and MIT (Boston, MA) using the Affymetrix Genome-Wide Human SNP Array 6.0. We thank the other investigators, the staff and the participants of the MESA study for their valuable contributions. A full list of participating MESA investigators and institutes can be found at http://www.mesa-nhlbi.org. L.F.B. was supported by R01 HL071739 for all measures of NAFLD in MESA. OOA studies are supported by grants and contracts from NIH, including U01 HL072515, U01 HL84756, U01 HL137181 and P30 DK72488. We acknowledge the MGI participants, Precision Health at the University of Michigan, the University of Michigan Medical School Central Biorepository and the University of Michigan Advanced Genomics Core for providing data and specimen storage, management, processing and distribution services. We also acknowledge the Center for Statistical Genetics in the Department of Biostatistics at the School of Public Health for genotype data curation, imputation and management in support of the research reported in this publication. COPDGene is supported by NHLBI (U01 HL089897 and U01 HL089856) as well as through contributions made to an industry advisory board comprised of AstraZeneca, Boehringer Ingelheim, GlaxoSmithKline, Novartis, Pfizer, Siemens and Sunovion. Liver fat measures in COPDGene were gathered under HL122464. Analyses in the UKBB were done under approved project 18120 (to E.K.S.). E.K.S., Y.C., A.K., X.D., A.O. and B.D.H. are supported by NIH (grants R01 DK106621 and R01 DK107904 to E.K.S.) and The University of Michigan Department of Internal Medicine. N.D.P. and E.K.S. are supported by NIH (grants R01 DK128871 to N.D.P. and E.K.S.; R01DK131787 to E.K.S.). V.L.C. was supported in part by an American Association for the Study of Liver Disease Clinical, Translational and Outcomes Research Award. We acknowledge the participants and investigators of the FinnGen study. The views expressed in this manuscript are those of the authors and do not necessarily represent the views of the NHLBI; the National Institutes of Health; the US Department of Health and Human Services; Framingham Heart Study or Boston University.
Author information
Authors and Affiliations
Contributions
E.K.S. led the conceptualization, methodology development and funding of the project. P.A.P., N.D.P. and E.K.S provided supervision of the project. E.K.S., A.K. and N.D.P. led project management. Analysis were conducted by Y.C. (lead), X.D. (lead), B.K., M.F.F., L.F.B., K.A.R., S.K.M., K.A.Y., X.G., A.V.S., A.K., A.O., N.D.P. and B.F.C. Study resources were provided by N.D.P., D.W.B., L.E.W., J.R.O., S.K.M., K.D.T., S.L.R.K., T.H.M., A.C., J.I.R., V.G., J.M.N., M.A.P., P.A.P., J.E.H., G.R.W. and E.K.S. Data curation was performed by M.A.A., M.J.B., J.J.C., J.G.T., Y.-D.I.C., G.E., B.D.H. and E.K.S; Y.C., X.D., N.D.P., P.A.P. and E.K.S participated in central results interpretation. Paper draft preparation and editing was performed by E.K.S. (lead), Y.C., A.K., V.L.C., X.D., A.O. and N.D.P. Final review: all authors. All authors had access to the study data and reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The Regents of the University of Michigan and E.K.S. have a pending patent on the use of systems and methods for analysis of samples associated with NAFLD and related conditions. V.L.C. received grant funding from KOWA and AstraZeneca. J.J.C. and Vanderbilt University Medical Center receive research funding from NIH, IBM Watson Health, GE Healthcare and Theratechnologies. G.R.W. is a cofounder and equity shareholder in Quantitative Imaging Solutions, a company that provides consulting services for image and data analytics. G.R.W.’s spouse works for Biogen. Grants or contracts from NIH, Department of Defense (DoD) and Boehringer Ingelheim made payments to G.R.W.’s institution. G.R.W. received consulting fees from Pulmonx, Vertex, Janssen Pharmaceuticals, Pieris Therapeutics and Intellia Therapeutics. G.R.W. also received payments from Pulmonx for participation on a Data Safety Monitoring Board or Advisory Board. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Stefano Romeo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1
GOLDPlus NAFLD measures meta-analysis study design.
Extended Data Fig. 2
European GOLDPlus NAFLD measures meta-analysis schematic.
Extended Data Fig. 3 Characteristics of GOLDPlus genome-wide significant variants in GOLD ancestry-based cohorts.
For each variant, the characteristics are shown for the GOLD ancestry-based analysis including: associated gene, NAFLD increasing effect allele (EA), effect allele frequency (EAF), effect/beta and 95% confidence interval (CI), Cochran’s Q heterogeneity I2 metric (HetSq) and heterogeneity P-value (HetPVal), EA P-value (P), and sample size (N). Results are for meta-analysis of GOLD European ancestry (red), African ancestry (blue), Hispanic ancestry (green), Chinese ancestry (purple), and all ancestries pooled (black). The estimates of the effect sizes (Beta) and 95% confidence interval in bidirectional testing within each ancestry and for all the ancestries combined were shown in the forest plots. The data underlying these plots are provided as Source Data.
Extended Data Fig. 4 Characteristics of GOLDPlus genome-wide significant variants in GOLD sex-specific cohorts.
For each variant, the characteristics are shown for the GOLD sex-specific analysis including: associated gene, NAFLD increasing effect allele (EA), effect allele frequency (EAF), effect/beta and 95% confidence interval (CI), Cochran’s Q heterogeneity I2 metric (HetSq) and heterogeneity P-value (HetPVal), EA P-value (P), and sample size (N). Results are for meta-analysis of GOLD cohort males (blue), females (red), and pooled sexes (black). The estimates of the effect sizes (Beta) and 95% confidence interval in bidirectional testing within each ancestry and for all the ancestries combined were shown in the forest plots. The data underlying these plots are provided as Source Data.
Extended Data Fig. 5 DEPICT analysis of biological enrichment of NAFLD associated variants.
Height of the bar represents the nominal −log10P-value of enrichment of GWAS associated genes with physiological systems, cells, and tissues. Dark orange shading represents statistical significance at false discovery rate (FDR) < 0.05. The data underlying these plots are provided as Source Data.
Extended Data Fig. 6 Two-sample Mendelian randomization analysis for casual associations between NAFLD associated variants and fibrosis/cirrhosis and esophageal varices.
a,b, Data represent the effect/beta and 95% confidence intervals for the inverse variance weighted (IVW) and MR-Egger analyses for (a) NAFLD exposure (GOLD cohort, n = 11 instruments) and K74:fibrosis/cirrhosis outcome (UKBB) (MR-Egger P-value = 1.88 × 10-3, IVW p-value = 8.65 × 10-5) and (b) NAFLD exposure (GOLD cohort, n = 11 instruments) and I85:esophageal varices outcome (UKBB) (MR-Egger P-value = 9.36 × 10-4, IVW P-value = 3.51 × 10-4). c,d, The crosshairs on the plots represent the effect and 95% confidence intervals for each SNP-NAFLD or SNP-outcome association for (c) NAFLD exposure (GOLD cohort, n = 10 instruments) and K74:fibrosis/cirrhosis outcome (UKBB) and (d) NAFLD exposure (GOLD cohort, n = 10 instruments) and I85:esophageal varices outcome (UKBB). The data underlying these plots are provided as Source Data.
Extended Data Fig. 7 Two-sample Mendelian randomization analysis for casual associations between BMI, waist circumference associated variants and NAFLD.
a,b, Data are presented as effect/beta and 95% confidence intervals for MR-Egger and inverse variance weighted (IVW) methods for (a) waist circumference GWAS (UKBB, n = 217 instruments) and GOLD cohort outcome (MR-Egger P-value = 3.6 × 10-2, IVW P-value = 3.71 × 10-4) and (b) BMI GWAS (UKBB, n = 293 instruments) and GOLD cohort outcome (MR-Egger P-value = 0.02, IVW P-value = 1.02 × 10-7). c,d, The crosshairs on the plots represent the effect/beta and 95% confidence intervals for each SNP-NAFLD or SNP-outcome association for (c) waist circumference GWAS (UKBB, n = 211 instruments) and GOLD cohort outcome and (d) BMI GWAS (UKBB, n = 283 instruments) and GOLD cohort outcome. The data underlying these plots are provided as Source Data.
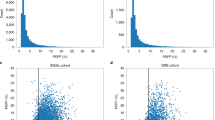
Extended Data Fig. 8 Convolutional neural network schematic for UKBB MRI liver imaging (PCC values).
Scatter plot of predicted UKBB MRI-PDFF values versus ‘true’ UKBB MRI-PDFF values (as determined by Perspectum Diagnostics). a,b, Pearson correlation coefficients (PCC) are shown for (a) gradient echo image protocol and (b) IDEAL image protocol. The data underlying these plots are provided as Source Data.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Tables 1–19.
Source data
Source Data Fig. 1
Source data to produce forest plots.
Source Data Fig. 2
Source data containing effect sizes for NAFLD-associated variants on human diseases and traits.
Source Data Fig. 3
Source data to produce forest plots.
Source Data Fig. 5
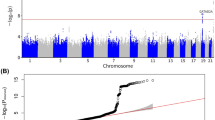
Source data to produce Manhattan plot.
Source Data Fig. 6
Source data to plot associations.
Source Data Extended Data Fig. 3
Source data to produce forest plots.
Source Data Extended Data Fig. 4
Source data to produce forest plots.
Source Data Extended Data Fig. 5
Source data that includes data from DEPICT analysis.
Source Data Extended Data Fig. 6
Source data that includes data from MR analysis.
Source Data Extended Data Fig. 7
Source data that includes data from MR analysis.
Source Data Extended Data Fig. 8
Source data that includes true and predicted PDFF.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Du, X., Kuppa, A. et al. Genome-wide association meta-analysis identifies 17 loci associated with nonalcoholic fatty liver disease. Nat Genet 55, 1640–1650 (2023). https://doi.org/10.1038/s41588-023-01497-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01497-6
This article is cited by
-
What are the common downstream molecular events between alcoholic and nonalcoholic fatty liver?
Lipids in Health and Disease (2024)
-
Comprehensive genetic study of the insulin resistance marker TG:HDL-C in the UK Biobank
Nature Genetics (2024)
-
Implications of the evolving knowledge of the genetic architecture of MASLD
Nature Reviews Gastroenterology & Hepatology (2024)
-
The contribution of genetics and epigenetics to MAFLD susceptibility
Hepatology International (2024)