Abstract
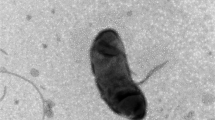
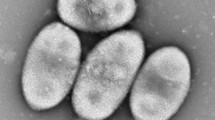
A gram-negative bacterium designated BN30T, which is motile with a polar flagellum, non-endospores forming, oxidase- and catalase-positive, was isolated from soybean root nodule. The organism is facultative anaerobic and surface-wrinkled rod. It can grow at 10–40 °C, pH 6–8 and 6 % (w/v) NaCl. BLASTn search based on 16S rRNA gene sequence revealed that the strain is closely related to Diaphorobacter aerolatus 8604S-37T, Alicycliphilus denitrificans K601T, Simplicispira limi EMB325T, Diaphorobacter nitroreducens NA10BT and Diaphorobacter oryzae RF3T, which all belonged to the family Comamonadaceae in class Betaproteobacteria. Phylogenetic analysis showed that the strain formed a firm clade with three Diaphorobacter species, being closest to Diaphorobacter aerolatus 8604S-37T with similarity of 98.64 %. DNA–DNA relatedness values between strain BN30T and five reference strains ranged from 11.5 to 35.9 %. All the results of phylogenetic analysis, chemotaxonomical data (predominant fatty acids are C16:0, sum feature 3, sum feature 8 and C17:0 cyclo; major polar lipids are diphosphatidylglycerol, phosphatidylethanolamine and phosphatidylglycerol; major quinone is Q-8; G+C of total DNA is 65.2 %), physiological and phenotypic results supported that BN30T represented a novel species within the genus Diaphorobacter. The name Diaphorobacter ruginosibacter was proposed, and the type strain is BN30T (=ACCC06116T = DSM 27467T).




Similar content being viewed by others
References
Borsodi AK, Makk J, Rusznyák A, Vajna B, Taba G, Márialigeti K (2007) Phenotypic characterization and molecular taxonomic studies on Bacillus and related isolates from Phragmites australis periphyton. Aquat Bot 86(3):243–252
Buck JD (1982) Nonstaining (KOH) method for determination of gram reactions of marine bacteria. Appl Environ Microb 44:992–993
De Lajudie P, Willems A, Nick G, Mohamed TS, Torck U, Filai-Maltouf A, Kersters K, Dreyfus B, Lindström K, Gillis M (1999) Agrobacterium bv. 1 strains isolated from nodules of tropical legumes. Syst Appl Microbiol 22(1):119–132
De Ley J, Cattoir H, Reynaerts A (1970) The quantitative measurement of DNA hybridization from renaturation rates. Eur J Biochem 12:133–142
Deng ZS, Zhao LF, Kong ZY, Yang WQ, Lindstrom K, Wang ET, Wei GH (2011) Diversity of endophytic bacteria within nodules of the Sphaerophysa salsula in different regions of Loess Plateau in China. FEMS Microbiol Ecol 76:463–475
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro RE (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Khan ST, Hiraishi A (2002) Diaphorobacter nitroreducens gen. nov., sp. nov., a poly(3-hydroxybutyrate)-degrading denitrifying bacterium isolated from activated sludge. J Gen Appl Microbiol 48:299–308
Kim O-S, Cho Y-J, Lee K, Yoon S-H, Kim M, Na H, Park S-C, Jeon YS, Lee J-H, Yi H (2012) Introducing EzTaxon-e: a prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. Int J Syst Evol Micrbiol 62(Pt 3):716–721
Kim SJ, Moon JY, Ahn JH, Weon HY, Hong SB, Seok SJ, Kwon SW (2014) Diaphorobacter aerolatus sp. nov., isolated from air, and emended description of the genus Diaphorobacter. Int J Syst Evol Micrbiol 64(2):513–517
Kuklinsky-Sobral J, Araujo WL, Mendes R, Geraldi IO, Pizzirani-Kleiner AA, Azevedo JL (2004) Isolation and characterization of soybean-associated bacteria and their potential for plant growth promotion. Environ Microb 6:1244–1251
Li JH, Wang ET, Chen WF, Chen WX (2008) Genetic diversity and potential for promotion of plant growth detected in nodule endophytic bacteria of soybean grown in Heilongjiang province of China. Soil Biol Biochem 40(1):238–246
Li L, Sinkko H, Montonen L, Wei G, Lindström K, Räsänen LA (2012) Biogeography of symbiotic and other endophytic bacteria isolated from medicinal Glycyrrhiza species in China. FEMS Microbiol Ecol 79:46–68
Lu S, Ryu SH, Chung BS, Chung YR, Park W, Jeon CO (2007) Simplicispira limi sp. nov., isolated from activated sludge. Int J Syst Evol Micrbiol 57(1):31–34
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J Mol Biol 3(2):208–218
Marmur J, Doty P (1962) Determination of the base composition of deoxyribonucleic acid from its thermal denaturation temperature. J Mol Biol 5:109–118
Mechichi T (2003) Alicycliphilus denitrificans gen. nov., sp. nov., a cyclohexanol-degrading, nitrate-reducing β-proteobacterium. Int J Syst Evol Micrbiol 53(1):147–152
Miller LT (1982) Single derivatization method for routine analysis of bacterial whole-cell fatty acid methyl esters, including hydroxy acids. J Clin Microbiol 16:584–586
Minnikin DE, Collins MD, Goodfellow M (1979) Fatty acid and polar lipid composition in the classification of Cellulomonas, Oerskovia and related taxa. J Appl Bacteriol 47(1):87–95
Pham VH, Park SJ, Roh Y, Roh DH, Rhee SK (2009) Diaphorobacter oryzae sp. nov., isolated from a thiosulfate-oxidizing enrichment culture. Int J Syst Evol Micrbiol 59:218–221
Powers EM, Latt TG (1977) Simplified 48-hour IMVic test: an agar plate method. Appl Environ Microb 34:274–279
Tindall BJ (1990a) A comparative study of the lipid composition of Halobacterium saccharovorum from various sources. Syst Appl Microbiol 13(2):128–130
Tindall BJ (1990b) Lipid composition of Halobacterium lacusprofundi. FEMS Microbiol Lett 66(1–3):199–202
Tittsler RP, Sandholzer LA (1936) The use of semi-solid agar for the detection of bacterial motility. J Bacteriol 31:575
Wang LL, Wang ET, Liu J, Li Y, Chen WX (2006) Endophytic occupation of root nodules and roots of Melilotus dentatus by Agrobacterium tumefaciens. Microb Ecol 52:436–443
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Zhang L, Xi L, Qiu D, Song L, Dai X, Ruan J, Huang Y (2013) Cellulomonas marina sp. nov., isolated from deep-sea water. Int J Syst Evol Microbiol 63:3014–3018
Acknowledgments
We are grateful to Professor Jean Euzéby for his helpful advice on the nomenclature; Professor Soon-Wo Kwon, for kindly supplying type strain Diaphorobacter aerolatus 8604S-37T; Professor Jie Gu and Ph. D Wei Sun for their supporting and help with the use of Biolog GN2 microplates. This research was financially supported by the 863 project of China (2012AA100402) and the National Science Foundation of China (31125007 and 31370142).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
The GenBank accession number for the 16S rRNA gene sequence of strain BN30T is KC825337. The strain had been deposited in Leibniz-Institut Deutsche Sammlung von Mikroorganismen und Zellkulturen Gmbh (DSMZ) and Agricultural Culture Collection of China (ACCC) under deposition number DSM27467 and ACCC06116. Reference strains Diaphorobacter nitroreducens NA10BT, Diaphorobacter oryzae RF3T, Simplicispira limi EMB325T and Alicycliphilus denitrificans K601T were bought from DSMZ; Diaphorobacter aerolatus 8604S-37T was supplied kindly by Professor Kwon (National Academy of Agricultural Science, Republic of Korea).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, X.L., Han, M.S., Xia, C.C. et al. Diaphorobacter ruginosibacter sp. nov., isolated from soybean root nodule, and emended description of the genus Diaphorobacter . Arch Microbiol 197, 683–692 (2015). https://doi.org/10.1007/s00203-015-1102-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-015-1102-7