Abstract
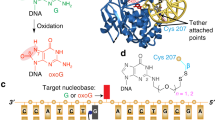
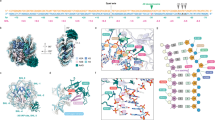
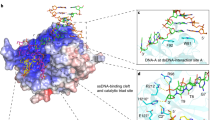
The Bacillus subtilis bacteriophages PBS-1 and PBS-2 protect their uracil-containing DNA by expressing an inhibitor protein (UGI) which inactivates the host uracil-DNA glycosylase (UDGase) base-excision repair enzyme. Also, PBS1/2 UGI efficiently inactivates UDGases from other biological sources, including the enzyme from herpes simplex virus type-1 (HSV-1). The crystal structure of the HSV-1 UDGase–PBS1 UGI complex at 2.7 Å reveals an α-β-α sandwich structure for UGI which interacts with conserved regions of UDGase involved in DNA binding, and directly mimics protein–DNA interactions observed in the UDGase–oligonucleotide complex. The inhibitor completely blocks access to the active site of UDGase, but makes no direct contact with the uracil-binding pocket itself.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Takahashi, I. & Marmur, J. Replacement of thymidylic acid by deoxyuridylic acid in the deoxyribonudeic acid of a transducng phage for Bacillus subtilis. Nature 197, 794–795.
Tomita, F. & Takahashi, I. A novel enzyme, dCTP deaminase, found in Bacillus subtilis infected with phage PBS1. Biochem. Biophys. Acta. 179, 18–27 (1969).
Price, A.R. & Fogt, S.M. Deoxythymidylate phosphohydrolase induced by bacteriophage PB52 during infection of Bacillus subtilis. J. biol. Chem. 248, 1372–1380 (1973).
Price, A.R. & Frato, J. Bacillus subtilis deoxyuridinetriphosphatase and its bacteriaphage PBS2 induced inhibitor. J. biol. Chem. 250, 8804–8811 (1975).
Hitzeman, R.A. & Price, A.R. Bacillus subtilis bacteriophage PBS2-induced DNA polymerase. J. biol. Chem. 253, 8518–8525 (1978).
Cone, R., Bonura, T. & Friedberg, E.C. Inhibitor of uracil-DNA glycosylase induced by bacterophage PBS2. Purification and preliminary characterisation, J. biol.Chem. 255, 10354–10358 (1980).
Savva, R. & Pearl, L.H. Cloning and expression of the uracil-DNA glycosylase inhibitor from bacteriophage PBS-1, and crystallisation of a uracil-DNA glycosylase-uracil-DNA glycosylase inhibitor complex. Proteins: Struct. Funct. Genet., in the press.
Wang, Z. and Mosbaugh, D.W. Uracil-DNA glycosylase inhibitor gene of bacteriophage PBS2 encodes a binding protein specific for uracil-DNA glycosylase. J. biol. Chem. 264, 1163–1171 (1989).
Wang, Z. & Mosbaugh, D.W. Uracil-DNA glycosylase inhibitor of bacteriophage PBS2: cloning and effects of expression of the inhibitor gene in Escherichia coli. J. Bacteriology. 170, 1082–1091 (1988).
Winters, T.A. & Williams, M.V. Use of the PBS uracil-DNA glycosylase inhibitor to differentiate the uracil-DNA glycosylase activities encoded by Herpes Simplex Virus types 1 and 2. J. virol. Meth. 29, 33–242 (1990).
Savva, R., McAuley-Hecht, K., Brown, T. & Pearl, L.H. The structural basis of specific uracil base-excision repair by uracil-DNA glycosylase. Nature 373, 487–493 (1995).
Mol, C.D. et al. Crystal structure and mutational analysis of human uracil-DNA glycosylase-structural basis for specificity and catalysis. Cell 80, 869–875 (1995).
Waksman, G. et al. Crystal structure of the phosphotyrosine recognition domain SH2 of v-src complexed with tyrosine-phosphorylated peptide. Nature 358, 646–653.
Sevcik, J., Hill, C.P., Dauter, Z., Wilson, K.S. Complex of ribonuclease from Streptomyces aureofaciens with 2′-GMP at 1.7Å resolution. Acta. Crystallogr. 49, 257–271 (1993).
Stubbs, M.T. et al The refined 2.4 Å X-ray crystal structure of recombinant human stefin B in complex with the cysteine proteinase papain: a novel type of proteinase inhibitor interaction. EMBO J. 9, 1939–1947.
Balasubramanian, S., Beger, R.D., Bennet, S.E., Mosbaugh, D.W. & Bolton, P.H. Secondary structure of uracil-DNA glycosylase inhibitor protein. J. biol. Chem. 270, 296–303 (1995).
Bennet, S.E., Schimerlik, M.I. & Mosbaugh, D.W. Kinetics of the uracil-DNA glycosylase/inhibitor protein association. Ung interaction with Ugi, nucleotides and uracil compounds. J. biol. Chem. 268, 26879–26885 (1993).
Savva, R. & Pearl, L.H. Crystallisation and preliminary X-ray analysis of the uracil-DNA glycosylase DNA repari enyzyme from herpes simplex virus type-1. J. molec. Biol. 234, 910–912 (1993).
Chayen, N.E., Shaw Stewart, P.D., Maeder, D.L. & Blow, D.M. An automated system for microbatch protein crystallization and screening. J. Appl. Crystallogr. 23, 287–302 (1990).
Leslie, A.G.W. Mosflm Users Guide, (MRC-LMB, Cambridge, UK, 1994).
Collaborative Computation Project, No. 4, Acta Cryst. D50, 760–763 (1994).
Navaza, J. AMoRE-an automated package for molecular replacement. Acta Crystallogr. A50, 157–163 (1994).
Brunger, A.T. X-PLOR Version 3.1 A system for X-ray crystallography and NMR (Yale University Press, New Haven, CT, USA, 1992).
Read, R.J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. 42, 140–149.
Laskowski, R.J., MacArthur, M.W., Moss, D.S. & Thornton, J.W. PROCHECK-a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–290.
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for builiding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Kraulis, P.J. MOLSCRIPT-a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Raster3D Version 20-a program for photorealistic molecular graphics. Acta Crystallogr. 50, 869–873 (1994).
Nicholls, A., Bharadwaj, R. & Honig, B. GRASP-graphical representation and analysis of surface properties. Biophys. J. 64, A116 (1993).
Wallace, A.C., Laskowski, R.A. & Thornton, J.M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions, Prot. Engng. 8, 127–134 (1995).
Engh, R.A. & Huber, R. Accurate bond and angle parameters for X-ray protein-structure refinement. Acta Crystallogr. A47, 392–400.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Savva, R., Pearl, L. Nucleotide mimicry in the crystal structure of the uracil-DNA glycosylase–uracil glycosylase inhibitor protein complex. Nat Struct Mol Biol 2, 752–757 (1995). https://doi.org/10.1038/nsb0995-752
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0995-752
This article is cited by
-
Antirestriction
Molecular Biology (2000)
-
The problem with pyrimidines
Nature Structural & Molecular Biology (1996)
-
A nucleotide-flipping mechanism from the structure of human uracil–DNA glycosylase bound to DNA
Nature (1996)