Abstract
Ulnar–mammary syndrome (UMS) is a rare autosomal-dominant disorder caused by mutations in TBX3. The condition is characterized by hypoplasia or aplasia of upper limbs on the ulnar side, mammary glands and nipples, and of apocrine glands in both sexes (MIM #181450). We report on a girl presenting with an UMS like phenotype, a dysmorphic facies, and mental retardation. Mutation analysis of TBX3 and G-banded chromosome analysis from lymphocytes were performed. We used microarray-based comparative genomic hybridization (array CGH) to investigate the patient's genomic DNA for submicroscopic aberrations. No mutation of the TBX3 gene was detected in our patient and chromosome analysis revealed a normal female karyotype (46,XX). Hybridization of a whole-genome tiling path array consisting of more than 36 000 BAC clones revealed an interstitial 1.28 Mb deletion within chromosomal band 12q24.21. The deleted region encompasses one known gene, TBX3. The deletion and haploinsufficiency of TBX3 was confirmed by fluorescence in situ hybridization using BAC clones representing the deletion on the BAC array. To our knowledge, this is the first description of TBX3 haploinsufficiency caused by a genomic deletion in a patient with UMS. We suggest that the UMS phenotype in conjunction with the characteristic facial changes and mental retardation observed in our patient is owing to the deletion of TBX3 and the involvement of neighbouring genes.
Similar content being viewed by others
Introduction
Ulnar–mammary syndrome (UMS) is a rare autosomal-dominant disorder (UMS, MIM #181450) with a characteristic upper limb malformation phenotype ranging from hypoplasia of the terminal phalanx of the fifth digit to aplasia of hand and upper limbs on the ulnar side.1 Abnormal development of mammary glands and nipples, teeth, genitalia, and of apocrine glands in both sexes are common clinical features in UMS patients.2, 3 Loss-of-function mutations in the TBX3 gene have been described to cause UMS.4, 5 TBX3 belongs to the T-box gene family, which encodes a large family of transcription factors sharing a DNA-binding domain called the T-box.4, 6, 7 Missense mutations, and nonsense mutations that result in impaired DNA binding as well as splice site mutations have been detected in UMS patients.4, 8 Single-base deletion and insertions (frameshift mutations) lead to a truncated protein lacking the T-box domain, which is incapable of binding DNA.9
Although our patient had typical features of UMS, no loss-of-function mutations in the TBX3 gene were detected. We therefore investigated the patient's genomic DNA for submicroscopic aberrations. Microarray-based comparative genomic hybridization (array CGH; Solinas-Toldo et al10and Pinkel et al11) has recently been implemented in molecular cytogenetics as a novel method to detect submicroscopic genomic aberrations.12, 13, 14 Higher resolution compared to conventional cytogenetic techniques makes array CGH a valuable tool in medical genetics. The use of tiling path arrays allows the detection of microdeletions or duplications as small as 100 kb.15 Using high-resolution array CGH, we detected a novel microdeletion on chromosome 12q24.21 encompassing the TBX3 gene.
Materials and methods
Clinical report
The female patient was the second child of unrelated healthy parents. Pregnancy and delivery were uneventful (Apgar scores 9/10). Anal atresia was detected after birth and surgically corrected. When seen by us at the age of 6 weeks, she presented with severe bilateral limb abnormalities. In addition, a dysmorphic facies and a short neck were noted (Figure 1). Family history was negative for similar conditions. At age 3.5 length and weight were at P3, and occipitofrontal circumference at P25. Dysmorphic features included capillary naevus flammeus on the central forehead (which faded with age), large eyebrows, flat nose, big mouth, small lips, slight pectus carinatum, and bilateral hypoplastic nipples (Figure 1a and b). Tooth eruption was delayed. Premature dental decay was noted. The bilateral limb abnormalities were more precisely described as mesomelic dysplasia of both arms and oligodactyly with hypoplastic fingers one, two, and three and contractures of the existing fingers on both hands as well as partial cutaneous syndactyly between the second and third fingers (Figure 1c, d and e). She walked at 2.5 years. A moderate mental retardation was present and speech development was delayed. X-ray examinations revealed bilateral distal hypoplasia of humeri, absent ulnae, and complete absence of fourth and fifth digits including the corresponding metacarpals and carpals (Figure 2). Bone age was delayed by 6 months. Radiographs of the chest, thoracal and lumbal spine, right leg, and foot were normal. Abdominal ultrasound examination, echocardiography, long-time electrocardiogram, and ocular examination disclosed no abnormalities, and hearing was normal.
(a) Facial appearance of the patient at the age of 6 weeks. Note capillary naevus flammeus on the forehead, large eyebrows, bulbous nose, and small lips. (b and c) Phenotype at the age of 3½ years. (b) Note inverted nipples, mesomelic dysplasia of the arms, bilateral absence of digit four and five; (c) mesomelic dysplasia of both arms, ulnar deviation of both hands; detailed picture of right (d) and left hand (e): absence of both fourth and fifth rays, hypoplastic fingers one, two, and three with flexion contractures of the middle interphalangeal joints of finger two and three on both hands, partial cutaneous syndactyly between second and third fingers.
Mutation analysis
Genomic DNA was prepared from peripheral lymphocytes as described previously. Exons 1–7 of TBX3 were PCR-amplified by means of previously published exon-flanking primers.4, 5 Sequencing was carried out at the lab of Mike Bamshad (Department of Pediatrics, Eccles Institute of Human Genetics, University of Utah).
Cytogenetics
Standard karyotyping of GTG-banded chromosomes from lymphocytes at 450 bands resolution was performed according to standard procedures. To exclude Fanconi anaemia cytogenetic investigation of induced chromosomal breakage in response to mitomycin C was performed on patient's lymphocytes.16
Fluorescence in situ hybridization
BAC clones RP11-636B16, RP4-601P9, RP11-33I06, RP11-297J16, RP11-722K12, and RP11-8A1 were obtained from the RZPD (Deutsches Resourcenzentrum für Genomforschung, Berlin, Germany). BAC DNA was fluorescently labelled using nick translation, and hybridized to metaphase spreads of the patient's lymphocytes using standard procedures. BAC clones were labelled with spectrum orange. YAC clone 933e11 mapping to 12p and cep12 (Vysis, Downers Grove, IL) labelled with spectrum green were used as control probes. Chromosomes were counterstained with DAPI. All colour images of hybridized metaphase spreads were evaluated using an epifluorescence microscope (Axioscope, Zeiss, Germany) fitted with different single band-pass filter sets for DAPI, Spectrum Green and Spectrum Orange fluorescence. The microscope is equipped with a cooled CCD camera (Hamamatsu) for image acquisition. Image analysis was performed using the ISIS analysis system (Metasystems, Altlussheim, Germany).
Microarray-based CGH
Array CGH was carried out using a 36k whole human genome tiling path BAC array consisting of the 1 Mb Sanger Clone set (kindly provided by Nigel Carter, Wellcome Trust Sanger Centre),17 a set of 390 subtelomeric clones (generated in the course of the EU initiative COSTB19: Molecular cytogenetics of solid tumours) and the human 32k Re-Array set (http://bacpac.chori.org/pHumanMinSet.htm; DNA kindly provided by Pieter de Jong).18, 19 Detailed protocols are available at the MPI webpage (http://www.molgen.mpg.de/~abt_rop/molecular_cytogenetics/ProtocolsEntry.html). The patient's DNA and a pooled reference DNA were fluorescently labelled with Cy5 and Cy3, respectively, using the Bioprime Array CGH labelling kit (Invitrogen, Carlsbad, CA, USA). Hybridization on a SlideBooster Hybridization Station (Implen, Munich, Germany) was performed as described in the protocol (refer to MPI webpage), except that DNA was not sonicated before labelling and 3 × 300 ng DNA were used. Images were scanned using GenePix 4100A and analysed using GenePix Pro 6.0 software (Axon Instruments, Foster City, CA, USA). Further analyses and visualization were performed with CGHPRO.20 35882 BACs were included in the analysis. Raw data were normalized by ‘Subgrid LOWESS’. The log 2 ratio of test to reference was calculated and plotted according to chromosomal position of the clones. Copy number gains and losses were determined by using a conservative threshold of 0.3 and −0.3, respectively. Aberrant signals including three or more neighbouring BAC clones were considered as genomic aberrations and were further evaluated by fluorescence in situ hybridization (FISH), unless they coincided with a published polymorphism as listed in the Database of Genomic Variants (http://projects.tcag.ca/variation/; version 13 December 2005).
Results
The patient described here presented with clinical features typical of UMS, that is, bilateral absent ulnae, complete absence of fourth and fifth digits (Figure 2), and anal atresia. In addition, facial dysmorphisms as well as mental retardation were noted, which have not been described as part of the UMS spectrum so far (Figure 1). The initially performed high-resolution karyotype was normal in the patient. Cytogenetic investigation of induced chromosomal breakage in response to mitomycin C yielded normal results. Mutation analysis by sequencing exons 1–7 of TBX3 gene detected no mutations. The additional features present in this patient suggested a submicroscopic rearrangement as the cause of the patient's condition. To test for such microdeletions, we performed array CGH analysis with the patient's genomic DNA using a whole-genome tiling path array. An interstitial deletion on chromosome 12q24.21 was detected (Figure 3). According to the array CGH data, the two breakpoints were located between RP11-636B16 and RP11-297J16 (proximal breakpoint) and RP11-8A1 and RP11-115H15 (distal breakpoint). The array data were mapped on the human genome sequence using the Ensembl genome browser (www.ensembl.org) to determine the deletion size and to identify candidate genes. The size of the deletion was determined to be ranging from 113.6 to 114.9 Mb. TBX3 is the only known gene located within the deleted region (positions 113.57–113.58 Mb). A second gene, THRAP2, is located distal to the deletion (positions 114.85–115.17 Mb). TBX5 is located proximal to the deletion (position 113.25–113.3 Mb).
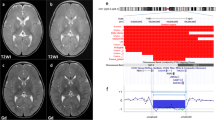
Array CGH profile. (a) Whole-genome view. Each spot in the profile represents a BAC clone. As BAC clones of the tiling path are overlapping, single clone aberrations were not considered true aberrations. For visualizing the content of low copy repeats in the ratio plots, each BAC clone was classified into one out of seven categories and colour-coded as described previously.20 (b) Detailed view of the interstitial deletion on chromosome band 12q24.21. The red and green lines indicate the log 2 ratio thresholds −0.3 (loss) and 0.3 (gain), respectively.
To confirm the array CGH data, we conducted FISH analysis using a set of BAC clones located at 12q24.21. BAC clone RP11-636B16 located at the proximal deletion breakpoint had an intermediate log 2 ratio (−0.21) in the array CGH experiment and showed normal FISH signals on chromosome 12 of patient's metaphases. BAC clone RP11–115H15 located at the distal deletion breakpoint showed two signals in the FISH experiment. As expected from the array data (log 2 ratio ⩽0.3), only one signal was detected on patient's metaphases for BAC clones RP4–601P9 (113.56 Mb), RP11–33I06 (113.57 Mb), RP11–297J16 (113.58 Mb), RP11–722K12 (114.2 Mb), and RP11–8A1 (114.67–114.84 Mb). An example is shown in Figure 4 (Supplementary information). Thus, the FISH results confirmed an interstitial deletion of approximately 1.28 Mb on chromosome 12 extending from 113.56 to 114.84 Mb and haploinsufficiency of the TBX3 gene. FISH on metaphase chromosomes of the patient's parents detected two signals on chromosome 12 for all probes investigated (data not shown), indicating that this deletion occurred de novo.
Discussion
The patient presented here shows a characteristic UMS phenotype, but has additional features that are not commonly considered to be part of the UMS phenotypic spectrum. To detect possible microdeletions, we analysed the genomic DNA by array CGH. Using a tiling path array, we were able to detect a de novo microdeletion of 1.28 Mb on chromosome 12q24.21 encompassing the TBX3 gene.
TBX3 is a member of the T-box gene family, which is involved in the specification of fore and hind limb identity. At least five T-box genes are expressed in the developing vertebrate limb, that is, TBX2, TBX3, TBX4, TBX5, and TBX15.21, 22 Loss-of-function mutations in TBX3 have been described to cause UMS, whereas TBX5 mutations are known to cause Holt–Oram syndrome (HOS).5, 23, 24 Bamshad et al4 investigated the genotype–phenotype correlation in the affected individuals of 10 families. No obvious clinical differences were detected between subgroups of patients with mutations in different TBX3 domains. Recently, Meneghini et al25 described an association between mutations affecting the T-domain of TBX3 and higher frequency of severe limb phenotypes. TBX3 is widely expressed in a variety of tissues and organs that are not affected in individuals with UMS or the TBX3 knockout mice. This suggests that different tissues may require different critical levels of normal TBX3 protein for morphogenesis or a regulation of TBX3 by tissue-specific cofactors. Alternatively, other genes from the T-box family as a consequence of functional redundancy may be able to compensate for reduced levels of normal TBX3 protein in other tissues than the forelimb. So far, no homozygous mutations have been observed in humans. Compared to patients with TBX3 point mutations, our patient presented with a rather severe manifestation of the syndrome. Bilateral symmetrical ulnar ray defects including the absence of the ulna as in the case presented here had previously been observed in TBX3 inactivation studies in the mouse and the chick,26 but have only been described in three out of 44 UMS patients from 10 families.4, 5
The characteristic facial appearance and the mental retardation observed in the patient described here are not typical for UMS. The large deletion is the most likely explanation for this broader phenotype. Although TBX3 is the only known gene within the deleted region, positional effects or other non-coding sequences of the deleted interval may have a regulatory role and may influence the expression of nearby genes such as TBX5, which is located at the proximal side of the deleted segment. Mutations in TBX5 cause HOS (OMIM 14290023, 24), a condition characterized by severe cardiac and skeletal abnormalities, in particular defects of the radial ray including triphalangeal thumb, hypoplastic/absent thumb/radius. Bilateral hypoplastic thumbs, hypoplastic fingers II and III, and hypoplastic bowed radii observed in our patient are radial ray defects, which might argue for a combined gene defect of TBX3 and TBX5. It is possible that the deletion of TBX3 and the genomic region upstream has a regulatory effect on the TBX5 gene. The THRAP2 (thyroid hormone receptor-associated protein 2, PROSIT240) gene is located distal to the deletion on 12q24.21. THRAP2 has been identified by two different groups while studying patients with balanced translocations.27, 28 The authors suggest that THRAP2 is a novel component of the thyroid hormone receptor-associated complex, which is involved in transcriptional regulation. The first case described by Muncke et al28 displays a congenital heart defect (transposition of the great arteries), delayed motor development, nearly absent speech, and mental retardation. The chromosomal breakpoint was located between the first and second exon of THRAP2. The second case was described to have a Noonan-like phenotype with normal intelligence and normal development (O Bartsch, personal communication). The breakpoint in this case is located 28 kb downstream of exon 1. The father had the same balanced translocation, but without clinical symptoms. The case presented here shares clinical features with the case described by Muncke et al,28 that is, mental retardation, delayed motor, and speech development. Analysis of THRAP2 expression detected high expression in skeletal muscle, heart, all subregions of the brain, and abundant expression in the foetal brain.28 The authors propose a possible role for THRAP2 in early heart and brain development. A disrupted or deregulated THRAP2 could therefore be a cause for developmental defects resulting in a retardation phenotype. This hypothesis is supported by three cases with a contiguous deletion of TBX5 and TBX3 with telomeric breakpoint positioned between 114.75 and 114.78 Mb (J Kohlhase, personal communication). All three carriers are of normal intelligence (J Kohlhase, personal communication), suggesting that the critical region for the retardation phenotype on 12q24.21 is between 114.78 and 114.84 Mb.
UMS associated with known TBX3 mutations has been interpreted as the result of haploinsufficiency for this gene. Our finding that a heterozygous deletion of the TBX3 gene also gives rise to UMS provides direct evidence for TBX3 haploinsufficiency as the mechanism causing this syndrome. The additional features observed in our patient clearly distinguish this phenotype from UMS caused by TBX3 mutations and argue for a positional effect and/or the involvement of other neighbouring genes.
References
Schinzel A : Ulnar–mammary syndrome. J Med Genet 1987; 24: 778–781.
Franceschini P, Vardeu M, Dalforno L et al: Possible relationship between ulnar–mammary syndrome and split hand with aplasia of the ulna syndrome. Am J Med Genet 1992; 44: 807–812.
Pallister P, Herrmann J, Opitz J : Studies of the malformation syndromes in man XXXXII: a pleiotropic dominant mutation affecting skeletal, sexual and apocrine-mammary development. Birth Defects Orig Artic Ser 1976; 12: 247–254.
Bamshad M, Le T, Watkins W et al: The spectrum of mutations in TBX3: genotype/phenotype relationship in ulnar–mammary syndrome. Am J Hum Genet 1999; 64: 1550–1562.
Bamshad M, Lin R, Law D et al: Mutations in human TBX3 alter limb, apocrine and genital development in ulnar–mammary syndrome. Nat Genet 1997; 16: 311–315.
Bollag R, Siegfried Z, Cebra-Thomas J, Garvey N, Davison E, Silver L : An ancient family of embryonically expressed mouse genes sharing a conserved protein with the T locus. Nat Genet 1994; 7: 383–389.
Herrmann B, Labeit S, Poustka A, King T, Lehrach H : Cloning of the T gene required in mesoderm formation in the mouse. Nature 1990; 343: 617–622.
Sasaki G, Ogata T, Ishii T, Hasegawa T, Sato S, Matsuo N : Novel mutation of TBX3 in a Japanese family with ulnar–mammary syndrome: implication for impaired sex development. Am J Med Genet 2002; 110: 365–369.
Wollnick B, Kayserili H, Uyguner O, Tukel T, Yuksel-Apak M : Haploinsufficiency of TBX3 causes ulnar–mammary syndrome in a large Turkish family. Ann Genet 2002; 45: 213–217.
Solinas-Toldo A, Lampel S, Stilgenbauer S et al: Matrix-based comparative genomic hybridization: biochips to screen for genomic imbalances. Genes, Chromosomes Cancer 1997; 20: 399–407.
Pinkel D, Segraves R, Sudar D et al: High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998; 20: 207–211.
Shaw-Smith C, Redon R, Rickman L et al: Microarray based comparative genomic hybridisation (array-CGH) detects submicroscopic chromosomal deletions and duplications in patients with learning disability/mental retardation and dysmorphic features. J Med Genet 2004; 41: 241–248.
Schoumans J, Anderlid B, Blennow E, Teh B, Nordenskjöld M : The performance of CGH array for the detection of cryptic constitutional chromosome imbalances. J Med Genet 2004; 41: 198–202.
Vissers L, Vries Bd, Osoegawa K et al: Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003; 73: 1261–1270.
Ishkanian A, Malloff C, Watson S et al: A tiling resolution DNA microarray with complete coverage of the human genome. Nat Genet 2004; 36: 299–303.
Tischkowitz MD, Hodgson SV : Fanconi anaemia. J Med Genet 2003; 40: 1–10.
Fiegler H, Carr P, Douglas EJ et al: DNA microarrays for comparative genomic hybridization based on DOP-PCR amplification of BAC and PAC clones. Genes Chromosomes Cancer 2003; 36: 361–374.
Krzywinski M, Bosdet I, Smailus D et al: A set of BAC clones spanning the human genome. Nucleic Acids Res 2004; 32: 3651–3660.
Osoegawa K, Mammoser AG, Wu C et al: A bacterial artificial chromosome library for sequencing the complete human genome. Genome Res 2001; 11: 483–496.
Chen W, Erdogan F, Ropers H, Lenzner S, Ullmann R : CGHPRO – a comprehensive data analysis tool for array CGH. BMC Bioinformatics 2005; 6: 85.
Gibson-Brown J, Agulnik S, Silver L, Niswander L, Papaioannou V : Involvement of T-box genes Tbx2–Tbx5 in vertebrate limb specification and development. Development 1998; 125: 2499–2509.
Agulnik S, Papaioannou V, Silver L : Cloning, mapping, and expression analysis of TBX15, a new member of the T-box family. Genomics 1998; 51: 68–75.
Basson C, Bachinsky D, Lin R et al: Mutations in human TBX5 [corrected] cause limb and cardiac malformations in Holt–Oram syndrome. Nat Genet 1997; 15: 30–35.
Li Q, Newbury-Ecob R, Terrett J et al: Holt–Oram syndrome is caused by mutations in TBX5, a member of the Brachyury (T) gene family. Nat Genet 1997; 15: 21–29.
Meneghini V, Odent S, Platonova N, Egeo A, Merlo G : Novel TBX3 mutation data in families with ulnar–mammary syndrome indicate a genotype–phenotype relationship: mutations that do not disrupt the T-domain are associated with less severe limb defects. Eur J Med Genet 2006; 49: 151–158.
Davenport T, Jerome-Majewska L, Papaioannou V : Mammary gland, limb and yolk sac defects in mice lacking Tbx3, the gene mutated in human ulnar–mammary syndrome. Development 2003; 130: 2263–2273.
Musante L, Bartsch O, Ropers H-H, Kalscheuer V : cDNA cloning and characterization of the human THRAP2 gene which maps to chromosome 12q24, and its mouse ortholog Thrap2. Gene 2004; 332: 119–127.
Muncke N, Jung C, Rüdiger H et al: Missense mutations and gene interruption in PROSIT240, a novel TRAP240-like gene, in patients with congenital heart defect (transposition of the great arteries). Circulation 2003; 108: 2843.
Acknowledgements
We thank Fabienne Trotier and Karen Stout-Weider for their excellent technical assistance in the array CGH and FISH experiments. We are also thankful to the patient and her family for their collaboration in this study. Additionally, we thank the BACPAC Resources Centre for providing the DNA of the human 32k Re-array set, the COST B19 Action for the clones of the subtelomeric set, the Mapping Core and Map Finishing groups of the Wellcome Trust Sanger Institute for initial clone supply and verification of the 1 Mb array and Claus Hultschig for spotting the arrays. The group of Mike Bamshad (Department of Pediatrics, Eccles Institute of Human Genetics, University of Utah) performed the TBX3 mutation analysis. This project was supported by the European Fund for Regional Development. The authors declare to have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Klopocki, E., Neumann, L., Tönnies, H. et al. Ulnar–mammary syndrome with dysmorphic facies and mental retardation caused by a novel 1.28 Mb deletion encompassing the TBX3 gene. Eur J Hum Genet 14, 1274–1279 (2006). https://doi.org/10.1038/sj.ejhg.5201696
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201696
Keywords
This article is cited by
-
The T-box transcription factor TBX3 drives proliferation by direct repression of the p21WAF1 cyclin-dependent kinase inhibitor
Cell Division (2016)
-
A multi-trait meta-analysis with imputed sequence variants reveals twelve QTL for mammary gland morphology in Fleckvieh cattle
Genetics Selection Evolution (2016)
-
A further case of the recurrent 15q24 microdeletion syndrome, detected by array CGH
European Journal of Pediatrics (2008)
-
Expanding the phenotype of alopecia–contractures–dwarfism mental retardation syndrome (ACD syndrome): description of an additional case and review of the literature
European Journal of Pediatrics (2008)